FAQ
My column is leaking buffer, what shall I do?
Leakage around connectors
| Possible causes | Suggested Remedy |
| Connectors not compatible with each other. | Check compatibility. |
| Connectors not compatible with solvents. | Check chemical resistance with the connector supplier. |
| Connectors poorly positioned or not tightened. | Check the connectors. |
| Gaskets worn out. | Gaskets lose flexibility with time and need to be replaced regularly. Inspect and replace if necessary and at least annually. |
Leaking tubing
| Possible causes | Suggested Remedy |
| Tubing not compatible with solvents. |
Check chemical resistance with the tubing supplier. Preventive action: Always check tubing solvent compatibility prior to packing or running the column. |
Leakage around end-pieces ( Not applicable for the pre-packed columns )
| Possible causes | Suggested Remedy |
| End-piece and O-rings not properly positioned with respect to the tube. | Disassemble the column and check the position of the end-piece and O-rings. Assemble the column according to instructions and perform leakage tests. |
| O-rings worn out | O-rings loose their flexibility with time and need to be replaced regularly. Disassemble the column and inspect the O-rings. Replace if necessary and at least annually. Assemble the column according to instructions and test for leakage. Preventive action: Replace O-rings when needed or at least annually. |
What is the pressure limit of my column?
Prepacked Columns
Please read
Prepacked chromatography columns for ÄKTAdesign systems
Other Columns
| Column |
Maximum pressure (bar)
|
| C 10, C 16 and C 26 |
1
|
| FineLINE Pilot 35 |
20
|
| HiScale |
20
|
| HR (High Resolution) 16 |
30
|
| SR 25 |
1
|
| Tricorn 5 |
100
|
| Tricorn 10 |
50
|
| XK 16 and XK 26 |
5
|
| XK 50 |
3
|
Can I run two columns in series to increase resolution or capacity?
Gel filtration chromatography
Running columns in series increases the resolution
Affinity chromatography, Hydrophobic interaction, Ion exchange chromatography
Reverse phase chromatography
Running columns in series increases the capacity but may have a bad impact on the resolution
due to increased dead volume.
Please note that back pressure may increase.
How would I get the best resolution of my column?
For stand alone columns please assemble the monitor to the column outlet.
If the column is connected to a system, connect the column as close as possible to the monitor.
My column has a gap between the packed bed and adaptor. Can I reuse the column?
| Possible causes | Suggested Remedy |
| Bed support damaged or incorrectly assembled allowing chromatography medium particles to leave the column. | Check the bed support and replace if necessary. Disassemble the column according to instructions. |
| Buffer conditions deviate with regard to temperature, conductivity, viscosity, content of organic solvent (reduces surface tension) or other factor. | Check the buffers and choose more suitable conditions. |
| Increased resistance to flow due to blocked bed support compressing the packed bed. | Clean or change the bed support. Disassemble the column and replace the support according to instructions. Preventive action: Pre-filter or centrifuge sample to avoid residues building up. |
| Poorly packed bed (not sufficiently compressed during packing). (Not applicable for prepacked columns) |
Evaluate the packing using recommended methods. Please be aware of that for small columns, less than 10 ml bed volume, the system dead volume has an impact on the column evaluation values. If the results are poor, refer to the symptom Poor packing evaluation in the troubleshooting section. |
The backpressure increases during operation
| Possible causes | Suggested Remedy |
| Auxiliary equipment such as manometers and pumps not working properly. | Check the function of all auxiliary equipment. Repair/replace if necessary. |
| Column is clogged. | Clean the column according to instructions. Choose the more rigorous cleaning protocol when available. See media and column instructions. |
| Bent tubing. | Check that the flow path is not restricted. |
| Buffer viscosity too high. | Check the viscosity of all buffers. Viscosity is a function of temperature. (Lower temperature gives higher viscosity.) Let low-temperature buffer reach operating temperature before starting the run. |
| Microbial growth in buffers. The buffer normally become opalescent due to microbial growth. |
Check buffers, especially those with phosphate, for microbial growth. Replace with fresh buffer if necessary. |
| Sample and collection vessels at different levels. | Adjust the vessels to approximately the same level. |
| The prefilter might be blocked. | Check the prefilter. Preventive action: Prefilters are not meant to substitute sample treatment. |
| Valve not fully open. | Check all valves. Open any that is not fully open. |
Find more causes and remedies in the troubleshooting section.
My column is clogged and/or discoloured. How can I clean my packed column?
Prepacked columns
Please clean the column according to instructions.
Other columns
Please follow the recommendations in the media instructions.
If the issue persists after cleaning you have to replace the column with a new one (prepacked columns) or repack the column with fresh medium.
How can I check the functionality of my column?
Leading and tailing peaks as defined according to the following figures:


| Leading peak | Tailing peak |
Experience shows that the best method of expressing the efficiency of a packed column is in terms of its height equivalent to a theoretical plate (HETP), reduced plate number (h) and its peak asymmetry factor (As).
Please be aware of that for small columns, less than 10 ml bed volume, the system dead volume has an impact on the column evaluation values.
How do I store my column?
Prepacked columns
Please store the column according to instructions.
Other columns
Please follow the recommendations in the media instructions.
My column contains air, what can I do?
| Possible causes | Suggested Remedy |
| Unclarified lysates may cause increased air bubble formation during purification. | An attached flow restrictor in the chromatography system can prevent this. If a flow restrictor is attached, it is important to change the pressure limit to 0.5 MPa (5 bar9 on the ÄKTAdesign system (where the column and the flow restrictor give a pressure of 0.3 MPa and 0.2 MPa respectively). |
| The column operates at room temperature after having been stored in a cold room. | Allow thermal equilibration before use. |
| - | Reverse the flow direction and pump well degassed water through the column. For recommended volumes and flow rates for your specific column, please refer to media and column instructions. |
More causes and remedies might be presented in the troubleshooting section.
What chemicals are compatible with my column?
Please see the chemical stability in the instructions for prepacked columns, empty columns and media.
How should I prepare my sample prior to loading the column?
Please prepare the sample according to instructions and handbooks
Prepacked column instructions
Please refer to relevant Instruction.
Other columns
Please refer to relevant Handbook.
My column has run dry. Can I reuse the column?
If the column still is free from bacterial growth it might be possible to reuse the column. It could be worth trying.
Remove the air by running liquid slowly from bottom to top of a vertical placed column at a pressure close to maximum column pressure. The size of air bubbles decreases with increased pressure and facilitate the bubbles escape from the column.
If you don't succeed in removing air from the column, the column must be replaced (prepacked columns) or repacked.
How much sample, in mg or ml of protein can I load onto my column?
Affinity Chromatography
- The binding capacity values listed in the selection guide below, are typical for the given species. However, there might be considerable deviations in binding capacity for different immunoglobulins derivated from same species, even if they are of the same subclass.
- Protein binding capacity is protein-to-protein dependent. The given capacities are to be considered as starting values.
- For optimal separation use approximately one fifth of the total binding capacity
For more information refer to:
Affinity chromatography columns and media selection guide
Glutathione Sepharose selection guide
Ni Sepharose and IMAC Sepharose selection guide
Desalting
- In group separation (desalting) the sample volumes can be up to 30% of the column volume.
- Regarding sample volumes for prepacked columns, please refer to Sample preparation for analysis of proteins, peptides and carbohydrates selection guide.
Gel filtration chromatography
- Choose a sample volume of < 0.5% of the column volume for media with average particle size in the range of 10-15 µm and < 5% of the volume for media with average particle size in the range of 30-100 µm.
- In group separation (desalting) the sample volumes can be up to 30% of the column volume.
- Regarding sample volumes for prepacked columns, please refer to selection guide below:
Sample preparation for analysis of proteins, peptides and carbohydrates selection guide.
Gel filtration columns and media selection guide and product profile selection guide.
Hydrophobic interaction chromatography
- The binding capacity values in the media and column instructions, there are examples and to be considered as starting values. Protein binding capacity is protein-to-protein dependent.
- For optimal separation use approximately one fifth of the total binding capacity.
Ion exchange chromatography
- The binding capacity values in the Ion exchange columns and media product profile, please refer to Ion exchange columns and media product profile selection guide, there are examples and to be considered as starting values. Protein binding capacity is protein-to-protein dependent.
- For optimal separation use approximately one fifth of the total binding capacity.
Reversed phase chromatography
- The binding capacity values in the media and column instructions, please refer to the related document, there are examples and to be considered as starting values. Protein binding capacity is protein-to-protein dependent.
- For optimal separation use approximately one fifth of the total binding capacity.
Accessories
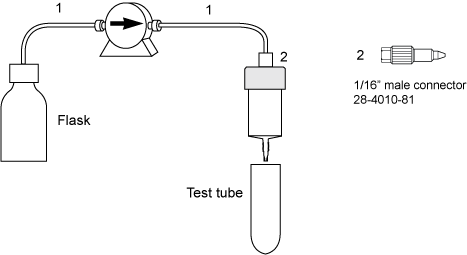
| # | Product Name | Product Code | Price | |
|---|---|---|---|---|
| 1 | Tubing cutter, for PEEK, EFTE, and FEP tubing i.d. 0.25, 0.5, 0.75, 1 and 1.6 mm | 18111246 | 99.55 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 0.75 mm, o.d. 1/16" | 18111253 | 96.26 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 0.5 mm, o.d. 1/16" | 18111368 | 73.48 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 1.0 mm, o.d. 1/16" | 18111583 | 85.34 USD |
Add to cart
|
| 1 | Tubing i.d. 0.25 mm, o.d. 1/16" | 18112095 | 66.24 USD |
Add to cart
|
| 2 | Fingertight connector 1/16" male, narrow | 28401081 | 108.65 USD |
Add to cart
|
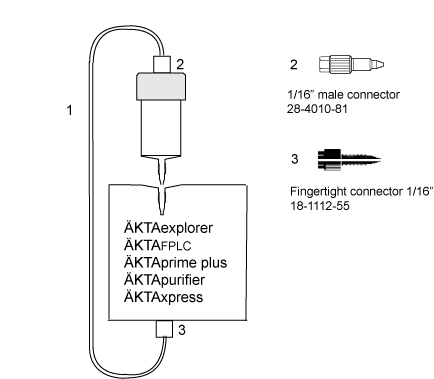
| # | Product Name | Product Code | Price | |
|---|---|---|---|---|
| 1 | Tubing cutter, for PEEK, EFTE, and FEP tubing i.d. 0.25, 0.5, 0.75, 1 and 1.6 mm | 18111246 | 99.55 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 0.75 mm, o.d. 1/16" | 18111253 | 96.26 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 0.5 mm, o.d. 1/16" | 18111368 | 73.48 USD |
Add to cart
|
| 1 | PEEK Tubing, 2 m, i.d. 1.0 mm, o.d. 1/16" | 18111583 | 85.34 USD |
Add to cart
|
| 1 | Tubing i.d. 0.25 mm, o.d. 1/16" | 18112095 | 66.24 USD |
Add to cart
|
| 2 | Fingertight connector 1/16" male, narrow | 28401081 | 108.65 USD |
Add to cart
|
| 3 | Fingertight Connector 1/16" Male for Tubing o.d. 1/16" | 18111255 | 123.16 USD |
Add to cart
|
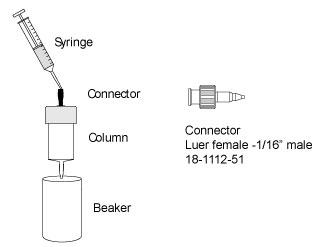
| # | Product Name | Product Code | Price | |
|---|---|---|---|---|
| 1 | Connector 1/16" Male/Luer Female | 18111251 | 109.24 USD |
Add to cart
|
Troubleshooting
Find solutions to product related issues. For unlisted issues please contact local Cytiva service representation.
Back pressure increases during operation
| Possible cause | Suggested remedy |
|---|---|
Auxiliary equipment such as manometers and pumps not working properly. |
Check the function of all auxiliary equipment. Repair/replace if necessary. |
Bent tubing. |
Check that the flow path is not restricted. |
Buffer viscosity too high. |
Check the viscosity of all buffers. Viscosity is a function of temperature. (Lower temperature gives higher viscosity.) Let low-temperature buffer reach operating temperature before starting the run. |
Microbial growth in buffers. |
Check buffers, especially those with phosphate, for microbial growth. Replace with fresh buffer if necessary. |
Sample and collection vessels at different levels. |
Adjust the vessels to approximately the same level. |
The prefilter might be blocked. |
Check the prefilter. |
Valve not fully open. |
Check all valves. Open any that is not fully open. |
Unusual column appearance
| Possible cause | Suggested remedy |
|---|---|
The column operates at room temperature after having been stored in a cold room. |
Allow thermal equilibration before use. |
| Possible cause | Suggested remedy |
|---|---|
Cleavage site sequence altered. |
Verify the presence of specific cleavage sites. |
Factor Xa is not properly activated. |
Activate factor Xa. Functional factor Xa requires activation of factor X with Russell´s viper venom. Activation conditions are a ratio of Russell´s viper venom to factor Xa of 1% in 8 mM Tris-HCI, 70 mM NaCl 8 mM CaCl2, pH 8.0. Incubate at 37oC for 5 min. Factor Xa from Cytiva has been preactivated by this procedure. |
Insufficient incubation time. |
Increase incubation time and enzyme concentration. For PreScission Protease, thrombin or factor Xa, increase the reaction time to 20 h or more if the tagged protein is not degraded by extensive incubation. The amount of enzymes can also be increased. |
Sample contains protease inhibitors. |
Ensure that proteas inhibitors are absent by performing a buffer exchange of the tagged protein against the cleavage buffer. |
The first amino acid after the factor Xa recognition sequence is Arg or Pro. |
Check the sequence of the fusion partner to be sure that the first three nucleotides, after the factor Xa recognition sequence do not code for Arg or Pro. |
The PreScission Protease, thrombin, or factor Xa to tagged protein ratios are incorrect. |
Check the amount of tagged protein in the digestion. Note that the capacity of GSTrap columns for GST is approximately 10 mg/ml medium in most purifications, however, the matrix is not saturated with tagged protein. Use the following ratios: |
| Possible cause | Suggested remedy |
|---|---|
Proteolysis occurred in host bacteria. |
Determine when the bands appear. Test to be certain that additional bands are not present prior to PreScission Protease, thombin or factor Xa cleavage. Such bands may be the result of proteolysis in the host bacteria. |
Tagged partner may contain recognition sequences for PreScission Protease, thrombin or factor Xa. |
Check the sequence of the cloned protein for protease recognition sequences. |
Unsatisfactory elution
| Possible cause | Suggested remedy |
|---|---|
Elution buffer is not fresh. |
Always use fresh elution buffer (reduced glutathione). |
Glutathione concentration is too low. |
Increase the concentration of glutathione in the elution buffer. The recommended concentration should be sufficient for most applications, but exceptions exist. Try 50 mM Tris-HCl, 20-40 mM reduced glutathione, pH 8.0 as elution buffer. |
Insufficient time for elution. |
Decrease the flow during elution. |
Insufficient volume of elution buffer. |
Increase the volume of elution buffer. Sometimes, especially after on-column cleavage of tagged protein, a large volume of buffer may be necessary to eluate the target protein. |
Ionic strength is too low |
Increase the ionic strength of the elution buffer. Adding 0.1-0.2 M NaCl to the elution buffer may improve results. |
Nonspecific hydrophobic interactions are interfering with elution. |
Add a nonionic detergent to the elution buffer. Nonspecific hydrophobic interactions may prevent solubilization and elution of target proteins from the column. Adding 0.1% Triton X-100 or 2% N-octylglucoside can significantly improve elution of some GST-tagged proteins. |
pH is too low. |
Increase the pH of the elution buffer. A low pH may limit elution from the column. Increasing the pH to 8-9 may improve elution without requiring an increase in the concentration of glutathione used for elution. |
| Possible cause | Suggested remedy |
|---|---|
Dead volume in chromatography systems is high. |
Minimize dead volume in the chromatography system by decreasing capillary length and dimensions between injector and detector. Bypass unused system components e.g. column valves from the flow path. |
Flow velocity too high. |
Run the separation at a lower flow velocity. This is especially important for adsorbents that bind several substances and where selectivity is low |
Too much sample has been loaded onto the column. |
Decreasing the sample load may improve the resolution significantly. |
| Possible cause | Suggested remedy |
|---|---|
Antibody used for detection in Western blotting may be cross-reacting with E. coli host proteins present in sample. |
Depending on the source of the anti-GST antibody, it may contain antibodies that react with various E. coli proteins that may be present in your tagged-protein sample. Cross-adsorb the antibody with an E. coli lysate to remove anti-E. coli antibodies fromt he preparation. Anti-GST antibody from Cytiva has been cross-adsorbed against E. coli proteins and tested for its lack of non-specific background binding in Western Blots. |
Chaperonins are co-purifying with GST-tagged protein |
Include an additional purification step. Additional bands may be caused by the co-purification of a variety of proteins known as chaperonins, which are involved in the correct folding of nascent proteins in E. coli . These include, but may not be limited to. |
Host strain is not protease deficient. |
Use a protease-deficient host. Multiple bands may be the result of proteolysis in the host bacteria. If this is the case, the use of a protease-deficient strain may be required (e.g. ion-or ompT) E. coli BL21 is provided with the pGEX vectors. This strain is ompT. |
Mr 70 000 protein is co-purifying with the GST-tagged protein. |
Mr 70 000 protein is probably a protein product of the E. coli gene dnaK. This protein is involved in protein folding in E. coli . It has been reported that this association can be disrupted by incubating the tagged protein in 50 mM Tris-HCl, 2 mM ATP. 10 mM MgSO2 pH 7.4 for 10 min. at 370C prior to loading on GSTrap 4 B. Alternatively, remove the DnaK protein by passing the tagged protein solution through ATP-agarose or by ion exchange. |
Over-sonication can cause copurification of host proteins with GST-tagged protein. |
Decrease sonication or change to another lysis method (e.g. French press). Cell disruption is apparent by partial clearing of the suspension and can be checked by microscopic examination. Adding lysozyme (0.1 volume of a 10 mg/ml lysozyme solution in 25 mM Tris-HCl, pH 8.0 prior to sonication may improve results. Avoid frothing as theis may denature the tagged protein. |
Partial degradation of tagged protein by proteases in lysate. |
Add a protease inhibitor. Adding 1 mM PM SF or Pefabloc SC to the lysis solution may improve results. |
| Possible cause | Suggested remedy |
|---|---|
Target protein not stable under the chosen conditions and partly degrades |
Find better ways to stabilize the protein, e.g. shorten the process time. |
The detergent has formed micelles with the protein, thereby increasing its size and changing its elution position. |
Reduce the concentration of detergent to below it´s critical micelle concentration (CMC ) value. |
| Possible cause | Suggested remedy |
|---|---|
Column not properly equilibrated. |
Check the pH and conductivity of the effluent before applying the sample. Continue to equilibrate with start buffer if necessary. |
Components in the sample displace the target molecule before elution starts. |
Reduce the concentration of detergent to below it´s critical micelle concentration (CMC ) value. |
| Possible cause | Suggested remedy |
|---|---|
The column not properly equilibrated |
Check the pH and conductivity of the effluent before applying the sample. Continue to equilibrate with start buffer if necessary. |
Column leakage
| Possible cause | Suggested remedy |
|---|---|
Connectors not compatible with each other. |
Check compatibility. |
Connectors not compatible with solvents. |
Check chemical resistance with the connector supplier. |
Connectors poorly positioned or not tightened. |
Check the connectors. |
Gaskets worn out. |
Gaskets lose flexibility with time and need to be replaced regularly. Inspect and replace if necessary and at least annually. |
| Possible cause | Suggested remedy |
|---|---|
Tubing not compatible with solvents. |
Check chemical resistance with the tubing supplier. |
Poor product recovery
| Possible cause | Suggested remedy |
|---|---|
GST-tagged protein denatured by sonication. Too extensive sonication can denature the tagged protein and prevent it binding to GSTrap columns. |
Use mild sonication conditions during cell lysis. Conditions for sonication must be empirically determined. |
| Possible cause | Suggested remedy |
|---|---|
Column is not equilibrated. |
Equilibrate the GSTrap column before use. Check that the GSTrap column has been equilibrated with a buffer pH 6.5 to 8.0 (e.g. PBS) before the cell lysate is applied. Binding of GST-tagged proteins to GSTrap columns is not efficient at pH less than 6.5 or greater than 8. |
Column needs cleaning. |
Clean the column according to instructions. If the GSTrap column has already been used several times, it may be necessary to replace the column with a new one |
Flow rate is too high. |
Decrease flow rate during sample load, see note in instructions |
Reducing agent is missing. |
Add DTT to a final concentration of 1-20 mM (GSTrap 4B) 1-10 mM (GSTrap FF and GSTrap HP) prior to cell lysis. This may significantly increase binding of GST-tagged proteins to GSTrap columns. |
Target protein may have altered conformation of GST, thereby reducing affinity for the medium. |
Test the binding of GST from the parental expression vector. Prepare a lysate of cells harboring the parental GST vector and check binding to the medium. If GST produced from the parental vector binds with high affinity, the fusion protein may have altered the conformation of GST, thereby reducing the affinity. Adequate results may be obtained by reducing the temperature used for binding to +4°C and by limiting column washing. |
Poor reproducibility
| Possible cause | Suggested remedy |
|---|---|
Continuous build-up of contaminants has altered the selectivity of the chromatography medium. |
Clean the chromatography medium according to instructions. |
| Possible cause | Suggested remedy |
|---|---|
Column not properly equilibrated |
Check the pH and the conductivity of the effluent before applying the sample. Continue to equilibrate with start buffer if necessary. |
Insufficient column regeneration. |
Prolong the regeneration. |
| Possible cause | Suggested remedy |
|---|---|
Protein properties change with concentrations |
Dilute or concentrate the sample to minimize effects. |
Proteins precipitate at high concentration. |
Reduce sample concentration and/or binding capacity. |
| Possible cause | Suggested remedy |
|---|---|
Column bleeding from previous run. |
Check and adjust your cleaning procedure. |
Column clogged with denatured proteins and/or lipids. |
Clean and regenerate the column and chromatography medium according to instructions. |
Incomplete equilibration of the column. |
Check pH and conductivity of the effluent before applying the sample. Continue to equilibrate if necessary. |
Incorrect pH and/or ionic strength of the solutions. |
Check pH and conductivity and adjust if necessary. Calibrate your conductivity and pH meters. |
Larger sample mass load applied compared with earlier runs. |
Keep mass of sample constant when repeating runs. (High proteins concentration can cause protein interaction, resulting in change of elution profile.) |
Sample volume is different from earlier runs. |
Resolution is dependent on the sample volume. Keep sample volume constant when repeating runs. |
| Possible cause | Suggested remedy |
|---|---|
Bound substances not removed during cleaning. |
Clean the chromatography medium according to instructions. |
Ligand partially degraded. |
Replace the chromatography medium. |
General advice to achieve good performance
Before using the column make sure that:
- Correct system has been selected in UNICORN System Control
- Correct wavelength has been set for UV/UPC monitor
- All tubing has been properly connected and tubing is not longer than needed
- All connectors are free from leakage, verified by passing a leakage test
- No tubing is folded or twisted
- Online filter, if used, is changed on a regular basis
- Correct buffers are used for the chosen columns and proteins
- All inlet tubing has been immersed in correct buffer solutions
- Enough buffer has been prepared
- Buffers have been equilibrated to the environment temperature
- Buffers/eluents have been degassed if necessary (e.g., in RPC runs)
- Suitable columns have been selected for the target proteins
- Column meets pressure requirements for selected medium
- Columns have been cleaned and prepared according to column instructions
- Samples have been clarified by centrifugation and/or filtration prior to sample loading
- Samples have been adjusted to binding buffer conditions
- Auto sampler (if used) has been prepared according to user manual
- The fraction collector has been filled with appropriate number of microtiter plates or tubes
- Appropriate arrangement for waste handling has been prepared
