It’s easy to think of Sanger sequencing as a technology of the past in this era of ever-increasing speed and scale. But Frederick Sanger’s DNA sequencing approach remains relevant even in today’s sequencing landscape—40 years after its introduction.
Let’s explore Sanger’s dideoxy chain termination method and how it continues to help scientists today.
What is Sanger sequencing?
Sanger sequencing is a method for determining the nucleotide sequence of DNA molecules. Developed by two-time Nobel laureate Frederick Sanger and his colleagues in 1977, it enabled an international collaboration of scientists to deliver the first human genome sequence.
By the mid-1970s, we had long known the basic structure of DNA and mechanism of transcription. But sequencing technology was in its infancy, and studying more genes and diseases in less time required a more efficient method to read DNA sequences.
The Sanger method relies on using dideoxy chain terminating nucleotides to produce sequence fragments of graduated lengths, each equal to the position of the base complementing the terminating nucleotide. Reading the DNA sequence involves visualizing these fragments and identifying the terminating nucleotides in order of fragment length (Fig. 1).
In the 1980s, developments in fluorescent detection, polymerase chain reaction (PCR), and electrophoresis improved the ease-of-use, speed, and sensitivity of Sanger sequencing. This automation-friendly approach led scientists to consider sequencing the human genome a real possibility—something only dreamt of a decade earlier!
Commercial Sanger sequencing equipment, such as the MegaBACE 4000 (Amersham International, now part of Cytiva) and PRISM 3700 (Applied Biosystems), were powerhouses of the Human Genome Project. Started in 1990, the project completed its objective—sequencing the human genome— in 2003, two years ahead of schedule.
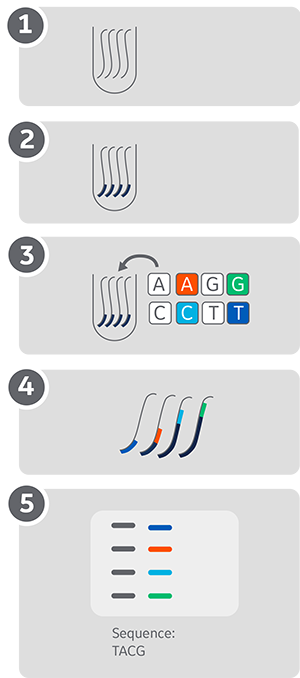
Fig. 1. Workflow for the Sanger method of DNA sequencing. Each dideoxynucleotide (ddNTP) is labeled using one of four different fluorescent dyes, enabling analysis on a single lane and automation.
1Starting with enough DNA sample, add a primer that binds to your region of interest.
2Add your labeled and unlabeled nucleotides. A small fraction (around 1%) of terminating fluorescently-labeled dideoxynucleotides (ddNTPs) enables fragment visualization. Each fluorescently labeled ddNTP carries a different fluorophore (red, yellow, blue, and pink).
3Synthesize to obtain your labeled fragments of varying lengths.
4Separate the fragments by capillary electrophoresis with single-base accuracy and find your original sequence.
Sanger in the era of next-generation sequencing
As the human genome project was ending, it was clear that to have a meaningful place in future healthcare, whole genome sequencing costs had to fall exponentially, and so the $1000 target was set.
New technologies emerged. Pyrosequencing and Illumina dye sequencing brought orders of magnitude increases in DNA sequencing speed and capacity. These massively parallel sequencing technologies, also dubbed next-generation sequencing (NGS), dominated the market for years.
But this did not mean the end of Sanger sequencing. The robustness and accuracy of Sanger sequencing, which can be as high as 99.9%, helped maintain its value over the years despite the availability of more modern alternatives.
One key area where Sanger continues to win out over NGS techniques is in small-scale sequencing applications. These include single-gene studies, routine sequencing for cloning and checking genotypes, and specialized and custom projects.
Whereas NGS provides huge capacity through parallel sequencing, this is less relevant in small-scale projects. Sanger sequencing gains an edge over NGS, with its higher accuracy, lower cost, and faster turnaround.
Sanger sequencing and NGS working together
Sanger sequencing can also complement NGS in, for example:
- Filling ‘gaps’ in NGS data in difficult-to-sequence areas and where coverage depth is low.
- Re-sequencing to confirm NGS results in small but critical sections of the genome.
- Validating new NGS approaches by analyzing a sample with both methods.
Filling the gaps with Sanger sequencing
When looking at large areas of the genome, scientists often plan their sequencing experiments based on achieving a minimum average coverage. However, even when the average coverage is high enough, there are often tricky areas where coverage is stubbornly low, such as GC-rich regions. Sanger sequencing continues to play a key role in filling in these gaps.
A good example is the detection of mutations in the CEBPA gene, which are a common cause for acute myeloid leukemia (AML). CEBPA is one of the key targets in cancer sequencing panels, but it also causes difficulties for NGS-based assays. High GC content (around 75%), presence of a trinucleotide repeat region, and frequent mutations in mononucleotide repeats all contribute to these difficulties.
As a result, CEBPA analysis with NGS produces poor amplicon coverage and technical issues in variant calling. These types of issues create an essential role for Sanger sequencing in supplementing NGS data and avoiding missed mutations.
Re-sequencing critical areas
Another reason to combine the two sequencing technologies is to confirm NGS data in critical areas. After sequencing large sections of DNA with NGS, re-checking small sections can increase the certainty of the presence or absence of a mutation.
In 2016, a study re-analyzed thousands of NGS variant calls with Sanger sequencing, investigating the relationship between NGS sensitivity, specificity, and false positive rate. It reviewed the data from a range of commercial NGS cancer panels that claimed 100% sensitivity in search of false positives. The researchers then also re-analyzed the data to give a revised sensitivity at which there were no false positives.
With the help of Sanger sequencing, the study found (at 100% sensitivity) a false-positive rate of 1.3%. Re-analysis to exclude false positives led to a revised sensitivity of 97.8%. These results indicate how NGS panels sometimes must strike a balance between sensitivity and specificity, whereas Sanger sequencing can generate more accurate answers.
Validation of NGS approaches
Using this approach of side-by-side testing, Sanger sequencing results can also provide a benchmark. Performing both methods on a few well-characterized samples can help validate the accuracy of an NGS approach before making it part of routine workflow.
These examples demonstrate how Sanger sequencing has kept its value to researchers over the decades. It’s an enduring technology that deserves a place in the toolkit of every scientist. Indeed, we might still be using it in another 40 years!
Find out more about optimizing your DNA sequencing workflows in other blogs. For support in any aspect of your workflow, contact our Scientific Support team or your local Cytiva representative.