We evaluated sanitization methods based on 1 M sodium hydroxide (NaOH) as sanitization agent using methodology previously developed for peracetic acid (PAA). We used an AxiChrom™ 300 column packed with pre-challenged Capto™ mixed-mode resin base matrix without coupled ligand. The chromatography resin was packed to a bed height of ~ 10 cm in a study using Pseudomonas aeruginosa as challenging organism.
Our main findings are:- The level of P. aeruginosa is reduced from 107 colony forming units (CFU)/mL of the challenging organism to levels of no detection throughout the whole packed bed including resin in the space between bed supports and tube.
- The method time for the procedure is under 2 h. This methodology also decreases the previous standard sanitization procedure time by 50% including securing no-flow zone sanitization.
The method described is also applicable for sanitization of a wide range of industrial chromatography resins tolerating 1 M NaOH such as Capto™, Sepharose™ Fast Flow and Sepharose™ High Performance. Our method enables sanitization either directly in the method for packing new resin or for packed columns already used in production.
Introduction
Microbial contaminations are found in many laboratory and production environments. Growing rapidly to large quantities under favorable conditions, these microorganisms can damage the function and impair the performance of chromatographic equipment and chromatography resin. Sodium hydroxide sanitization provides an effective means to purge downstream bioprocess resins from microbial contaminants. NaOH cleaning-in-place is the widely accepted method of sanitization employed by biopharmaceutical manufacturers to avoid the excessive costs and delays of a bioburden event in their downstream bioprocess.
We have previously described a sanitization method using AxiChrom™ 300 large-scale bioprocess columns with the sporicidal agent, peracetic acid (PAA) [1]. Here, we describe a microbial challenge test like the one used in the PAA study but with a concentration of 1 M NaOH instead to mimic typical sanitization practices employed by the biopharma industry for a broad range of resins. The final version of our method with resin and chemically dependent hold times is highly efficient as an integral part of the packing procedure.
Microbial challenge test
Microbial challenge tests introduce a high concentration of a predetermined microbial organism into the equipment, after which the challenged surfaces and resin are treated with an antimicrobial agent. After a specified time, the numbers of surviving organisms are counted.
Here, we performed challenge tests on Capto™ mixed mode resin without coupled ligand packed in an AxiChrom™ 300 column with gram-negative P. aeruginosa as challenging organism. The resin was pre-challenged with 107 CFU/mL of P. aeruginosa. After sanitization with 1 M NaOH, the process liquids and the dismantled column parts and resin were sampled for remaining microorganisms at predetermined sites.
Test setup and criteria for acceptance
Criteria of acceptance were aligned with standards set out by the United Sates Pharmacopeia (USP) and discussions with pharmaceutical collaborators in the bioprocessing industry, that is:
- Challenging organism > 106 CFU/mL in the homogenous resin slurry applied.
- Liquid samples after finishing the sanitization procedure—including a 5 to 6 day (d) temporary storage state (“clean hold”) in 20% ethanol—must have a maximum concentration of 10 CFU/10 mL of contaminating organism. Additionally:
- A maximum of 10% of samples taken during column disassembly may be contaminated with microorganisms other than the challenging microorganism.
The detailed methods are described in the Materials and methods section at the end of this document.
Results and discussion
We developed a sanitization method based for AxiChrom™ 300 column packed with high-flow agarose resin without coupled ligand by running a study using P. aeruginosa as challenging organism. The concentration of the challenging organism in the prepared pre-contaminated resin suspension was as required above 106 CFU/mL.
Effective removal of the P. aeruginosa contaminating organism
Table 1 shows that no challenging organism was discovered in any of the 60 samples taken during the study and we can therefore conclude that the acceptance criteria set at the beginning of the study were entirely met.
Table 1. A summary of the total number of samples found contaminated during this study
| Contaminating organism | Number of CFU |
| Challenging organism (P. aeruginosa) | 0 |
| Other organisms | 6 |
| Total number of samples analyzed 1 | 60 |
1 Samples that are not referred to in the acceptance criteria including negative or positive controls are excluded (five samples in total).
Other sources of contamination
Contaminants other than the challenging organism were found on three sample points during swabbing: a resin sample and a point at the interface between outer bed support steel ring and net were contaminated with Gram-positive rods (spore formers) that we could not identify. Despite these findings, this study met the acceptance criteria in terms of effective removal of P. aeruginosa and was therefore approved.
Control samples in Table 2 indicate that materials and procedures did not pose a risk of compromising the studies. Furthermore, we observed that the initial rinse with 1.9 CV of filtered purified water has good reductive effect of more than log 5 on the concentration of the challenging organism (data not shown).
Table 2. Contaminant microorganisms detected at different steps and different sampling points described in the study
| Sample | Unit | Day 0 | Day 6 |
| A 100 mM PAA before application in the rinse channel | CFU/50 mL | 0 | – |
| B Slurry solution with challenging organism (before application) | CFU/mL | 1.8 ×107 | – |
| C Chasing with 50 mM NaCl (before application) | CFU/50 mL | 0 | – |
| D Effluent from finished consolidation | CFU/mL | 7.9 × 106 | – |
| E Filtered rinse water before application | CFU/50 mL | 1* | – |
| F Flowthrough after 1.9 CV of filtered rinse water | CFU/mL | 20 | – |
| G 20 mM PAA solution (before application) | CFU/50 mL | 0 | – |
| H Flowthrough after 2 CV of filtered rinse water | CFU/50 mL | 0 | – |
| I Flowthrough after rinse/equilibration with 2 CV of 20% ethanol (Day 0) | CFU/50 mL | 4* | – |
| J Flowthrough of resin valve in the bottom inlet (Day 0) | CFU/50 mL | 12* | – |
| K Liquid from the rinse channel (100 mM PAA). 20% ethanol is used to push out the PAA (Day 0) | CFU/50 mL | 0 | – |
| L Filtered 20% ethanol before application (Day 6) | CFU/50 mL | – | 0 |
| M Flowthrough taken after 0.6 CV after rinse/equilibration with 20% ethanol (Day 6) | CFU/50 mL | – | 0 |
| N Flowthrough taken after 1.2 CV after rinse/equilibration with 20% ethanol (Day 6) | CFU/50 mL | – | 0 |
| O Flowthrough of resin valve in the bottom inlet (20 % ethanol) (Day 6) | CFU/50 mL | – | 0 |
| P Liquid from the rinse channel (20% ethanol) (Day 6) | CFU/50 mL | – | 0 |
* airborne organisms from higher-than-normal air levels in the lab
Conclusions
- We efficiently sanitized AxiChrom™ 300 column packed with Capto™ mixed-mode resin base matrix using 1 M NaOH in under 2 h.
- The count of of P. aeruginosa was reduced from 107 CFU/mL to an acceptable level when analyzed after a 6 d period of clean-hold storage.
- Method automation ensures robustness and repeatability of sanitization and 1 M NaOH can be recommended for sanitization of a wide range of industrial chromatography resins based on high-flow agarose, such as Capto™ mixed-mode chromatography resins.
Materials and methods
All parts in contact with the process flow including the resin were precleaned, challenged with 107 CFU/mL P. aeruginosa, and sanitized with 1 M NaOH, and evaluated. After sanitization, the process liquid was sampled, and the column was dismantled.
Samples of predetermined sites on the column as well as the packed resin were collected for analysis of remaining P. aeruginosa. Flowthrough samples were collected from the process chamber during the run and from the rinse channel, process chamber, and resin valve after finishing the sanitization method.
Finally, the cleaned column was filled with storage solution (20% ethanol) and left in temporary storage state (clean hold). Additional flowthrough samples were collected after 6 d of clean hold and analyzed for contaminating microorganisms before the column was disassembled for swabbing.
Preparation of the column, ÄKTAprocess™ system
An AxiChrom™ 300 column with new and unused acrylic tube and stainless-steel bed supports was the starting point for our study. The bottom and top bed support net (10 µm) were of the latest design, which includes the 273 × 2.62 mm EPDM O-ring (29165936) for sealing off the interface between the top plate and the adapter. All O-rings in contact with the process stream and directly or indirectly connected to a sampling point were replaced with new ones.
We performed the sanitization study, including cleaning of column parts, in a hygienic lab with controlled airflow. All column parts that could be disassembled (column lid, tube, bottom, and adapter backing plate excluded) were soaked in 1 M NaOH solution for 24 h, then rinsed with autoclaved purified water before assembling. The remainder of the column (column lid, bottom, and the adapter backing plate) was sprayed with 70% ethanol, while the column tube was wiped with 20% ethanol before assembly.
Before soaking the column parts in 1 M NaOH for 24 h, all parts were scrubbed or wiped with a 2% mild washing up liquid detergent solution, except for small parts such as screws and nuts. Stainless-steel bed supports were cleaned in an ultrasonic bath with 1 M NaOH for 2 × 15 min at 40°C.
The polytetrafluoroethylene (PTFE) thread tape on all screws including the fasteners for bed supports was replaced after which the screws with new tape were autoclaved for 30 min at 121°C. When possible, column parts were assembled in a laminar airflow (LAF) fume hood. During assembly the parts were sprayed with Ecolab™ Klercide™ 70/30 denatured ethanol (Ecolab USA, Inc).
ÄKTAprocess™ liquid chromatography system was precleaned according to application note CY13228-07May20-AN.
Column pre-sanitization and packing with inoculated resin
The column was primed using the Intelligent Packing method for AxiChrom™ columns.
After priming, the column was: 1. Filled with 1 M NaOH with the adapter at 40 cm and stored overnight. 2. The adapter was moved down to starting position, 1 cm from the bottom bed support at which the column was equilibrated with purified water to pH neutral.
Before the column was packed, we filled the flush channel (the area between the upper and lower scraper seals) with 100 mM PAA for sanitization using a syringe. The column was packed with Capto™ mixed-mode resin base matrix using 50 mM NaCl as packing solution inoculated with the challenging organism.
The homogenous slurry of the infected resin (107 CFU/mL P. aeruginosa) was drawn into the column by raising the adapter, initially at 300 cm/h and at the end around 100 cm/h since all the resin was chased into the column. Chasing was performed by pouring 50 mM NaCl into the slurry tank while the adapter was still moving upwards in the column to completely empty the tank from infected slurry, chasing it into the column with packing buffer. The resin valve was then closed, and the valve and tubing were rinsed free from resin using water and then 1 M NaOH.
The packing was started with the bottom mobile phase open while the adapter was driven downwards at 60 cm/h to a target bed height of 9.7 cm.
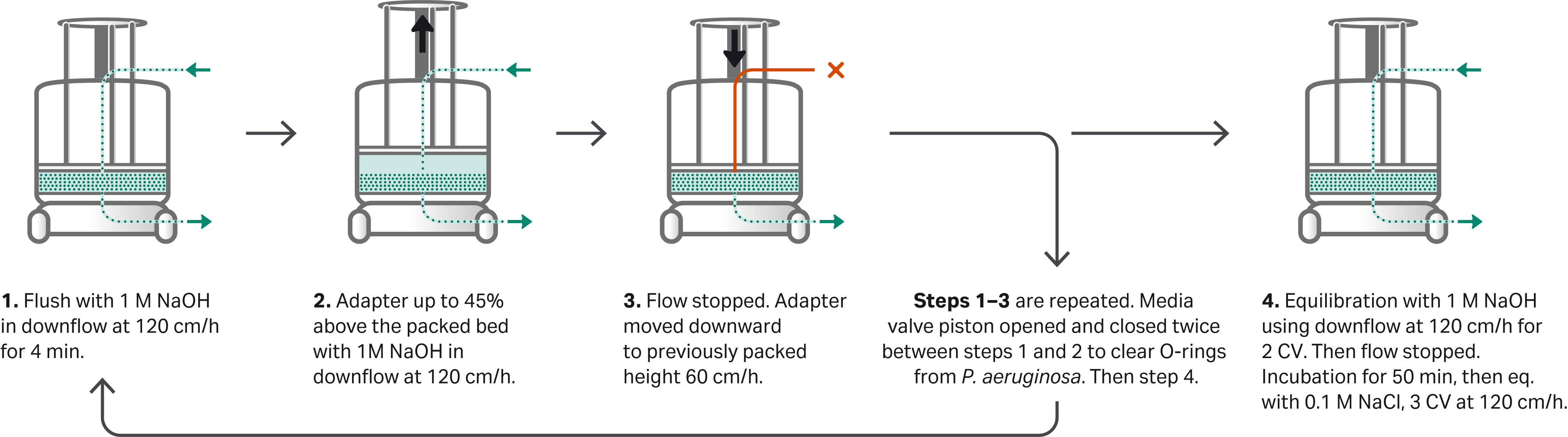
The sanitization steps with inoculated resin are summarized in in the schematic diagram, see Figure 1.
Fig 1. Schematic diagram of the sanitization procedure covering important stages in the process. The step-by-step protocol is described in the section Column sanitization procedure below.
The changes made to the method from the previously described PAA study were adapted in a proof of concept study and resulted in adapter hold times of 2 × 15 min to let the resin fall and create turbulence and solution exchange in the no-flow zone. This hold time is dependent on resin and different sanitization solutions and requires method adaptations in terms of hold time adjustment.
Column sanitization procedure
The following sanitization method was run automatically by a programmed UNICORN™ method and the automated sanitization steps are summarized earlier in Figure 1:
- Sanitization of the packed bed was initiated by a post-infection rinse of the packed bed with 2 CV of sterile purified water in a downflow at 60 cm/h.
- A rinse with 1 M NaOH in downflow at 120 cm/h for 0.83 CV was performed, which forced the solution into the packed bed (step 1 in Fig 1).
- The adapter was moved upwards at 120 cm/h to 13.7 cm while 1 M NaOH was pushed through the bed downflow at 170 cm/h (step 2 in Fig 1) and kept in at position for 15 min. (step 3 in Fig 1).
- Flow was stopped and instantly the adapter moved downwards to 10.3 cm cm at 60 cm/h, pushing excess liquid out through the bottom mobile phase (step 3 in Fig 1).
- A second rinse with 1 M NaOH in downflow at 120 cm/h for 0.83 CV was performed, which forced the NaOH solution further into the packed bed.
- The flow was again stopped, and the resin valve piston was open and closed twice within 6 s to clear the resin valve O-rings from trapped challenging organisms, from the contaminated resin filling.
- The adapter was moved upwards a second time at 120 cm/h to 13.7 cm while 1 M NaOH was pushed through the bed downflow at 170 cm/h, again keeping the adapter raised and with continuous flowing for 15 min to let all the resin in the groove fall.
- The flow was stopped, and the adapter instantly moved downwards a second time to 10.3 cm at 60 cm/h pushing excess liquid out through the bottom mobile phase.
- A rinse was performed by running a downflow with 1 M NaOH at 120 cm/h (step 4 in Fig 1).
- The flow was stopped and the NaOH-treated bed was incubated for 50 min to reach a resin contact time of at least 60 min (step 4 in Fig 1).
- Finally, the packed bed was equilibrated with 0.1 M NaCl by running a downflow at 120 cm/h for 3 CV. To facilitate liquid sampling, the flow was reduced to 60 cm/h at the end of this step (step 4 in Fig 1).
Microbial sampling
Liquid sampling points were collected:
(1) after the consolidation phase of the packing before step 1 in Figure 1;
(2) at a rinse with 2 CV of water after packing and before step 1;
(3) after equilibration with 2 CV of purified water in step 4.
After sanitization, the packed bed and resin valve were equilibrated with 20% ethanol and liquid samples were collected after 2 CV for the packed bed and 3 L for the resin valve. The 100 mM PAA in the rinse channel was rinsed out with ~ 150 mL 20% ethanol until pH increased and then a liquid sample was collected.
The column was then incubated with 20% ethanol for a 6-d clean hold while connected to the ÄKTAprocess™ system. Before any liquid samples were collected after the clean hold, the system was flushed with and equilibrated with 20% ethanol to minimize the risk of contaminants from the system.
A liquid sample of the ethanol in the resin valve was collected and then 20% ethanol was run with downflow at 60 cm/h through the packed bed and liquid samples were collected after 0.6 CV and 1.2 CV. A liquid sample was also collected from the top mobile phase of the filtered 20% ethanol as a control. The liquid in the flush channel was also collected.
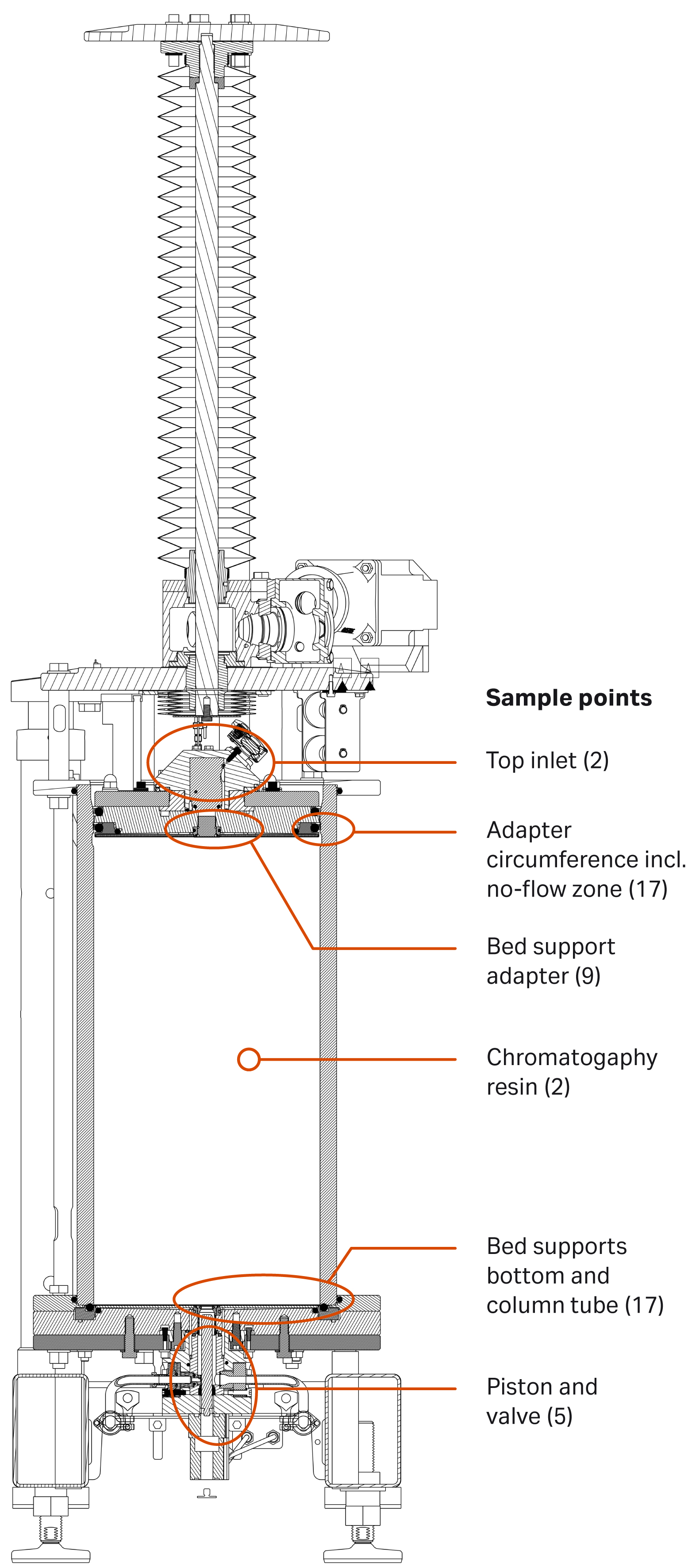
After the clean hold, column disassembly was performed, and microbial swab were taken from the predetermined hardware sites as shown in Figure 2.
Fig 2. Predetermined swab sampling points of the AxiChrom™ 300 column (numbers in brackets indicate the number of samples analyzed).
Microbial sampling was performed by one of the following methods:
Direct filtration
Sample solutions (50 mL) were collected in sterile tubes and then filtered through 0.45 mm cellulose nitrate membrane filters. We incubated the filters on agar plates at 30°C to 35°C for 5 d after which the plates were inspected for CFU.
Swabbing
Surface samples were taken with swabs. The swab was inserted into the tube containing the isotonic swab rinse solution and vortexed for a minimum of 20 s. The solutions including the swabs were poured into Petri dishes and mixed with 30 mL of temperature controlled molten agar. After solidification, plates were incubated at 30°C to 35°C for 5 d after which the plates were inspected for CFUs.
Agar plate
A sample of the chromatography resin was taken after sanitization and mixed with 30 mL of molten agar.
One gram of drained resin was aseptically transferred into a sterile container. We added the molten agar aseptically to the container and mixed with the resin to homogenously suspend it. The suspension was transferred to and allowed to solidify in Petri dishes, which were incubated at 30°C to 35°C for 5 d and subsequently inspected for CFU.
Viable count
Samples of challenging organism suspensions were diluted in series in 0.9% NaCl. Samples from the diluted suspensions were plated on agar plates and incubated at 30°C to 35°C for 1 to 2 d after which the plates were inspected for CFU. The concentration of challenging organism was determined in the sampled suspensions.
TR29681727
CY20698