This white paper is part of our Biacore™ SPR applications series.
In this series, we explore how Biacore™ SPR technology supports a wide range of applications—from membrane protein analysis to fragment screening, antibody-drug conjugates (ADCs), and beyond. Each article highlights real-world data and uses case studies to help researchers unlock the full potential of SPR.
Key takeaways:
- Antibody-drug conjugates (ADCs) have emerged as potential therapeutics due to their specificity, which enhances the therapeutic index and reduces side effects.
- Biacore™ SPR systems can be used as an essential tool in the development of therapeutic ADCs, enabling the evaluation of antibody and antigen interactions and binding kinetics, including how these are affected by conjugation, pH, linker chemistry, and drug-antibody ratios.
- Biacore™ SPR technology enables real-time assessment of the kinetics and affinity of anti-drug antibodies and ADCs, including weak-affinity and transient reactions, to produce accurate, reproducible data that can accelerate therapeutic ADC development.
Introduction to antibody-drug conjugates
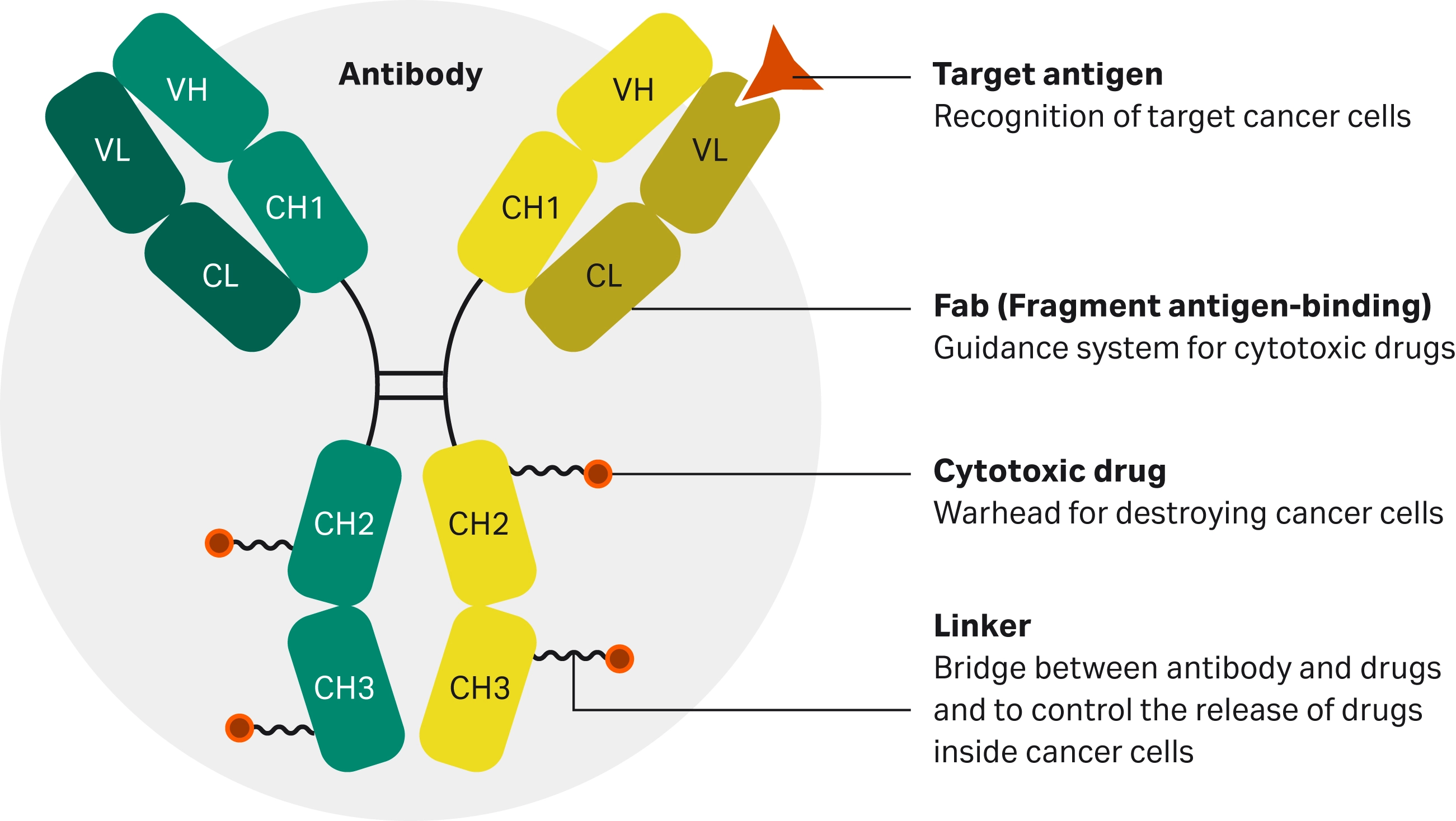
ADCs have great potential for use in a wide variety of diseases, particularly cancer (1). They are comprised of a monoclonal antibody that targets specific antigens, a cytotoxic payload designed to induce cell death, and a chemical linker designed to facilitate drug release once inside the target cells (2) (Fig 1).
Fig 1. Diagram of the core constituents forming an ADC (2).
ADCs are increasing in popularity as a potential method to treat diseases, with fifteen ADCs currently approved by the FDA as of April 2025 (3), and over one hundred ADC candidates involved in clinical trials (1). Previously, diseases like cancer have been treated with systemic therapies, such as chemotherapy. Due to their systemic nature, these cytotoxic agents attack both cancer cells and healthy cells indiscriminately, resulting in severe side effects and a limit to how toxic the agent can be. ADCs' revolutionary function is in the specificity with which they target pathologies, enhancing the therapeutic index and reducing side effects. Their method of action involves (2):
- The antibody region of the ADC recognizes and binds to antigens specific to or overexpressed in pathology, such as HER2 on breast cancer cells, and facilitates ADC internalization.
- The linker remains stable in circulation and releases the cytotoxic drug only inside the target cell, triggered by intracellular conditions or enzymatic cleavage.
- The cytotoxic payload exerts its cytotoxicity after the internalization of the ADC into the target cell.
Analytical hurdles in ADC development
Although ADCs have the potential to transform the way that diseases are treated, multiple analytical hurdles remain in ADC development. One key challenge is assessing how antibody-antigen binding specificity may be affected by drug conjugation, linker type, or drug-to-antibody ratio (DAR), where a reduction in binding affinity may affect ADC specificity and undermine its therapeutic effectiveness (4). When carrying out these binding kinetics and affinity assessments, the use of labels can affect binding kinetics, necessitating a label-free technique (5). A further consideration in ADC development is how physiological conditions (pH, temperature, ionic strength, etc.) influence ADC binding, to ensure that the ADC is optimized to its target environment (4). In addition, immunogenicity and the development of anti-drug antibodies (ADAs) are major concerns in therapeutic ADC development, as ADAs can neutralize ADCs, alter their pharmacokinetics, and increase safety risks. Therefore, it is essential that the interactions between all ADAs and ADCs are thoroughly evaluated. However, this can be challenging if ADAs only have weak affinity or when there is a high concentration of ADCs in the sample (6, 7). As a result, specialized analytical tools, such as SPR, are essential for generating robust and reproducible data to support ADC development.
ADC drug discovery and development pipeline
The ADC development pipeline involves selecting a tumor-specific antibody, linking it to a cytotoxic payload via a cleavable linker, and then developing the complete ADC for targeted cancer therapy. Surface plasmon resonance (SPR) applications within this pipeline include target identification, antibody characterization (measuring binding affinity and kinetics of target protein binding), epitope binning, and pre-clinical drug screening to accelerate lead selection and optimization of ADC candidates, by providing real-time, label-free data on molecular interactions (Table 1).
Table 1. Summary of ADC development pipeline
| ADC development pipeline | Objective | SPR relevance |
|---|---|---|
| Target selection | Identify antigens or receptors that are overexpressed on cancer cells but minimally present on healthy tissue. | SPR enables real-time analysis of antigen-antibody interactions, helping to validate target specificity and binding strength early in the discovery phase. |
| Antibody development | Engineer or select monoclonal antibodies (mAbs) with high specificity and affinity for the target antigen. | SPR enables high-throughput screening to identify clones expressing the desired mAb, followed by detailed characterization of binding kinetics and affinity, guiding the selection of optimal antibody candidates. |
| Payload selection | Choose a potent cytotoxic payload capable of killing cancer cells upon internalization. | While SPR is not used to assess cytotoxicity directly, it can help evaluate whether payload conjugation affects antibody binding performance. |
| Linker development |
Design a stable linker that connects the antibody to the payload and ensures controlled release within the target cell. |
SPR can assess the impact of linker chemistry on antibody-antigen binding, providing indirect insights into linker stability and ADC integrity. |
| ADC assembly |
Synthesize the complete ADC molecule with a consistent drug-to-antibody ratio (DAR) and site-specific conjugation. |
SPR can verify that conjugation does not compromise antigen binding and can be used to compare different conjugation strategies. |
| In vitro and in vivo testing |
Evaluate potency, toxicity, stability, and efficacy in cell-based and animal models. |
SPR supports preclinical analysis by providing safety and potency attributes across ADC variants. |
| Clinical trials |
Assess safety and efficacy in human patients. |
SPR contributes to clinical development by supporting comparability studies, stability testing, and potency assays under GMP conditions. |
Biacore™ surface plasmon resonance technology overview
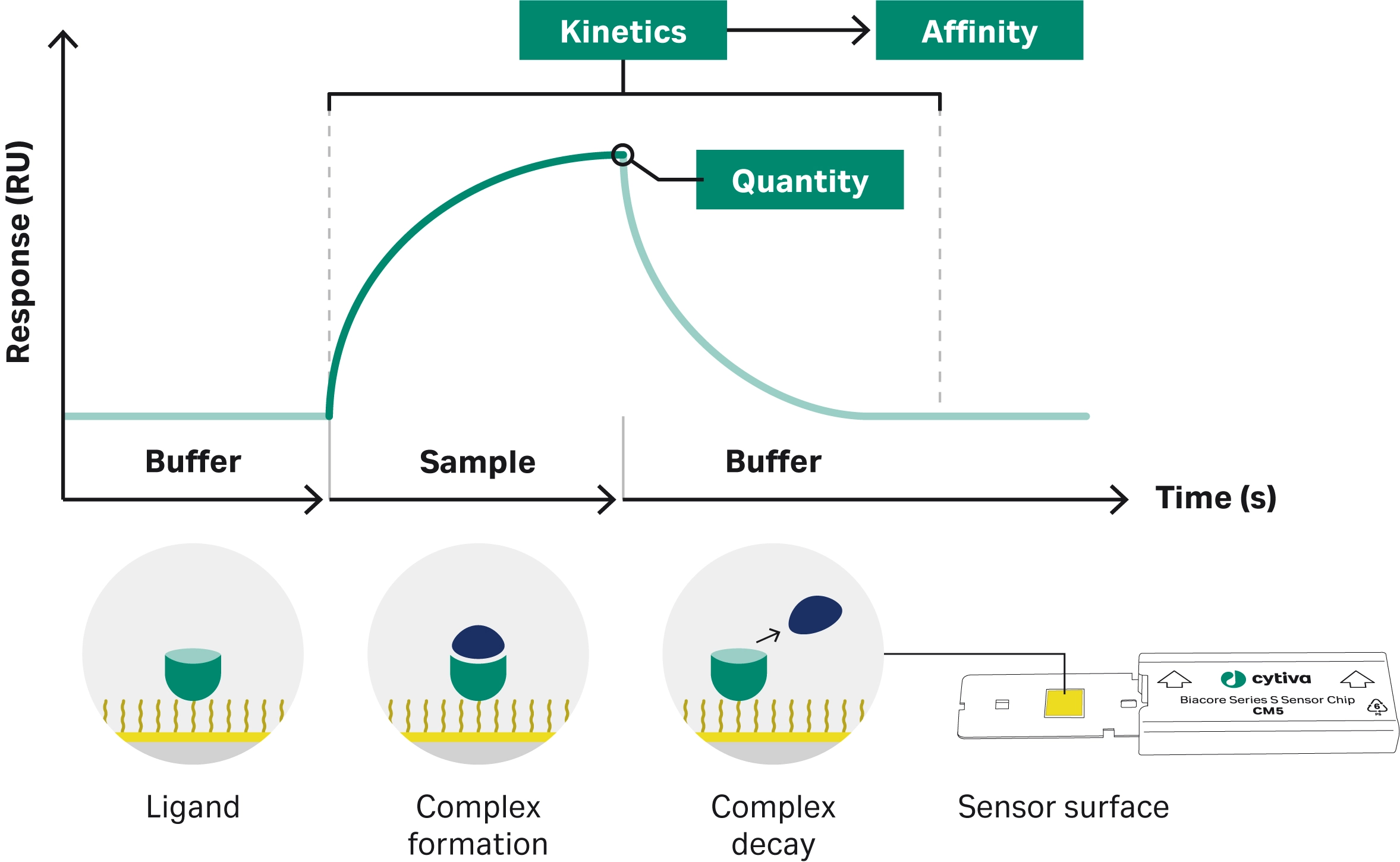
Surface plasmon resonance (SPR) is an optical technique that detects molecular interactions in real time without the need for labeling. In Biacore™ SPR systems, one of the interaction partners is attached to an SPR sensor surface, while the other flows over the sensor surface. Binding events generate a response proportional to the bound mass on the surface, with sensitivity down to picograms. The resulting plot, known as a sensorgram (Fig 2), represents binding response over time, and allows detailed analysis of interaction characteristics including kinetics (ka, kd), affinity (KD), and active concentration. Biacore™ SPR systems support a wide range of other applications, such as yes/no binding studies, specificity testing, potency determination, epitope binning, and comparability assessments, making it a powerful tool for confident decision-making in ADC characterization workflows.
Fig 2. Biacore™ SPR system with a sensorgram display showing real-time association and dissociation of molecular interactions.
How does Biacore™ SPR help in ADC characterization?
Biacore™ SPR technology is increasingly being used in the development of therapeutic ADCs to analyze binding interactions between the ADC and its target.
Evaluating antibody-antigen binding kinetics
The assessment of antibody-antigen binding kinetics is crucial for ADC development. For an ADC to be successful, the antibody must bind to the antigen with sufficient binding affinity and specificity to ensure that the therapeutic effect occurs at the target cell.
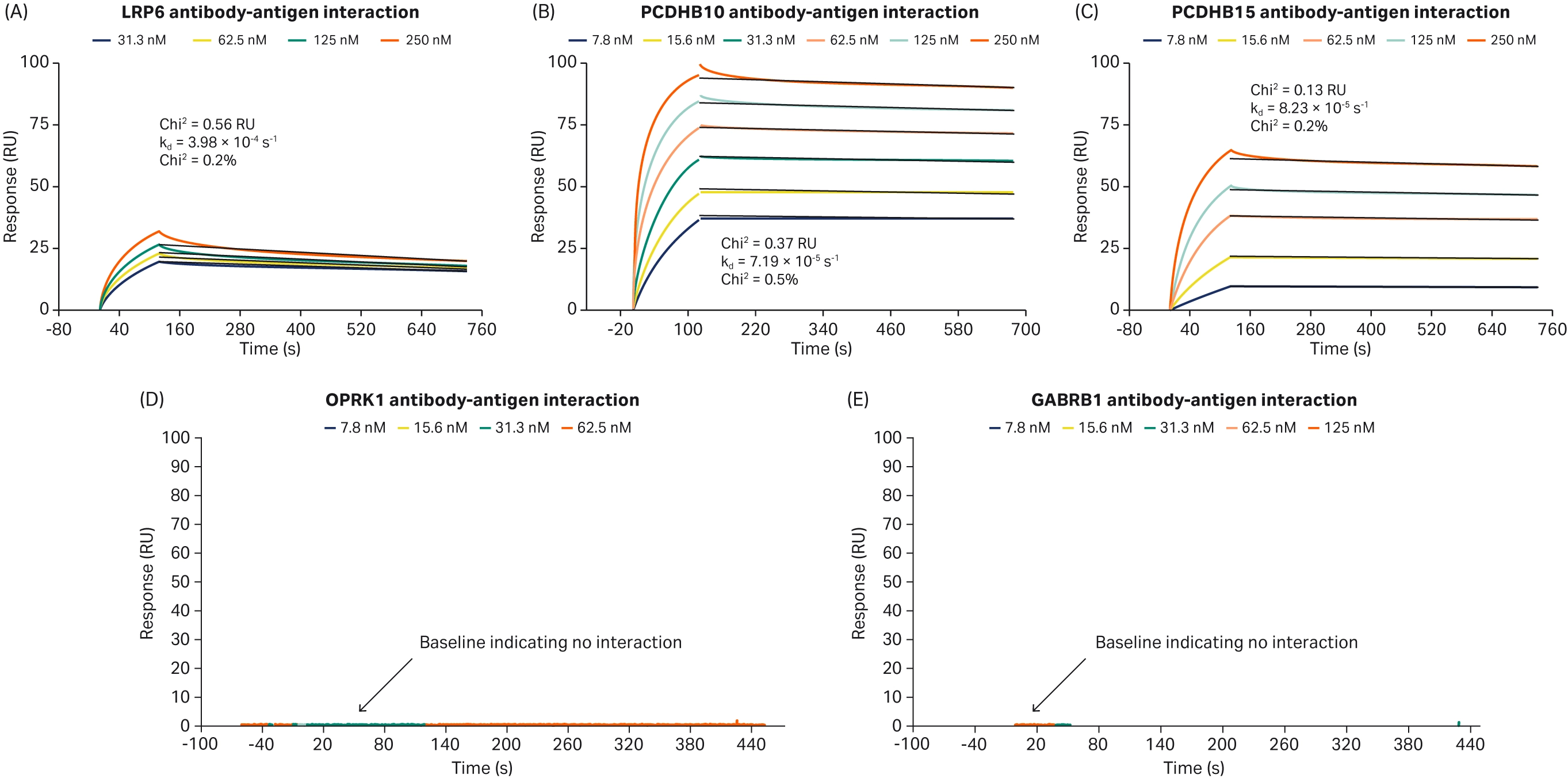
The use of Biacore™ SPR technology in assessing the binding kinetics between potential antibodies and the target antigen is common (8-10). In a study investigating the development of ADCs targeting gynecological cancers, Healey et al. used Biacore™ SPR analysis to assess the binding characteristics of four potential antibodies against rRAGE as the antigen (4). They demonstrated that only one antibody showed sufficient binding affinity (Fig 3). Similarly, Onyido et al. used Biacore™ SPR technology to assess the binding of five potential antibodies to their respective target proteins (11). The target proteins were injected over antibodies directly coupled onto the SPR sensor chip CM5 via NHS–EDC chemistry. The resultant binding profiles, using the 1:1 dissociation model, revealed that binding of the recombinant proteins to their respective antibody was very stable, with off-rates (kd) ranging from 4 × 10−4 to 8 × 10−5 s−1. Three of the antibodies were demonstrated to have stable binding, whereas the remaining two antibodies did not show any binding (Fig 4). These studies both demonstrate how SPR can be applied early in ADC development to screen and make informed decisions on potential antibody candidates.
Fig 3. SPR sensorgrams of the four potential antibodies (RBG01-04) binding to the rRAGE antigen (4).
Fig 4. SPR sensorgrams of three potential antibodies with (A-C) stable binding and (D-E) no binding (11).
Assessing the effect of conjugation on binding kinetics
ADC production requires the conjugation of the cytotoxic payload to the antibody via a linker. However, conjugation has the potential to alter antibody binding to the target protein. As a result, it is essential that the effect of conjugation on the antibody-antigen interaction and binding kinetics is assessed during ADC development.
Acchione et al. investigated the effect of conjugation on the antigen-binding ability and stability of ADCs (12). Biacore™ SPR technology was used to determine the binding kinetics of each potential antibody, and this was compared to each antibody’s corresponding ADC. The binding kinetics were not found to be significantly different for either of the ADCs assessed, enabling further development and testing of the ADCs. In a similar study, Cardillo et al. also compared the ADC-binding with the parent antibody binding, demonstrating that conjugation did not adversely affect binding to the antigen, and in fact enhanced the binding affinity compared to the parental antibody (13). Chen et al. used Biacore™ SPR technology to investigate whether conjugation affected the antigen-binding of their potential biosimilar, finding minimal influence of conjugation (14). Healey et al. also used Biacore™ SPR analysis to assess two of their potential ADCs before and after conjugation (4). They found a four-fold reduction in the binding affinity to the antigen for one ADC (Fig 5. A-B), and no discernible effect on binding affinity for the second (Fig 5. C-D).
Fig 5. SPR sensorgrams comparing the binding kinetics of unconjugated versus conjugated antibodies (4).
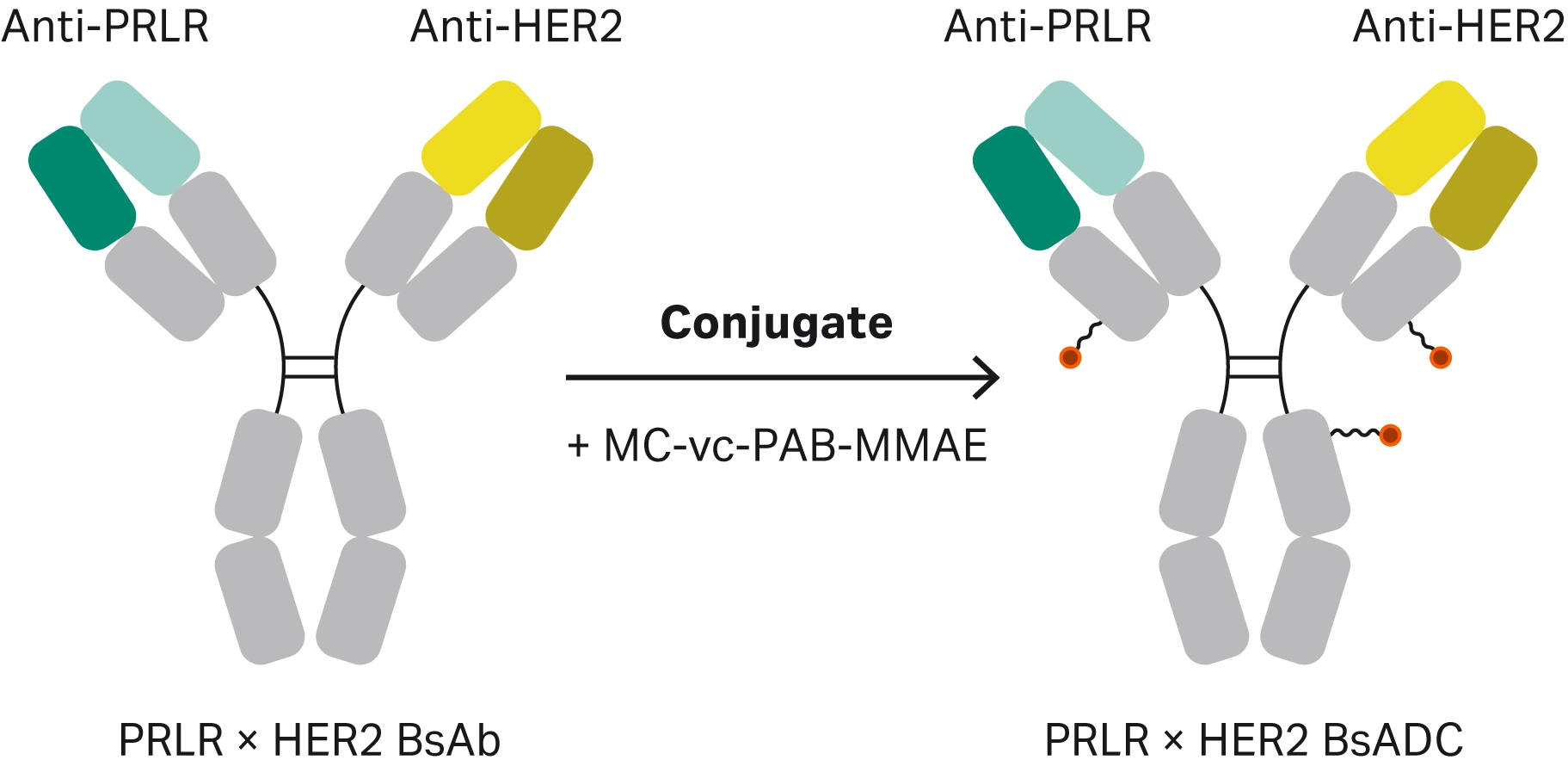
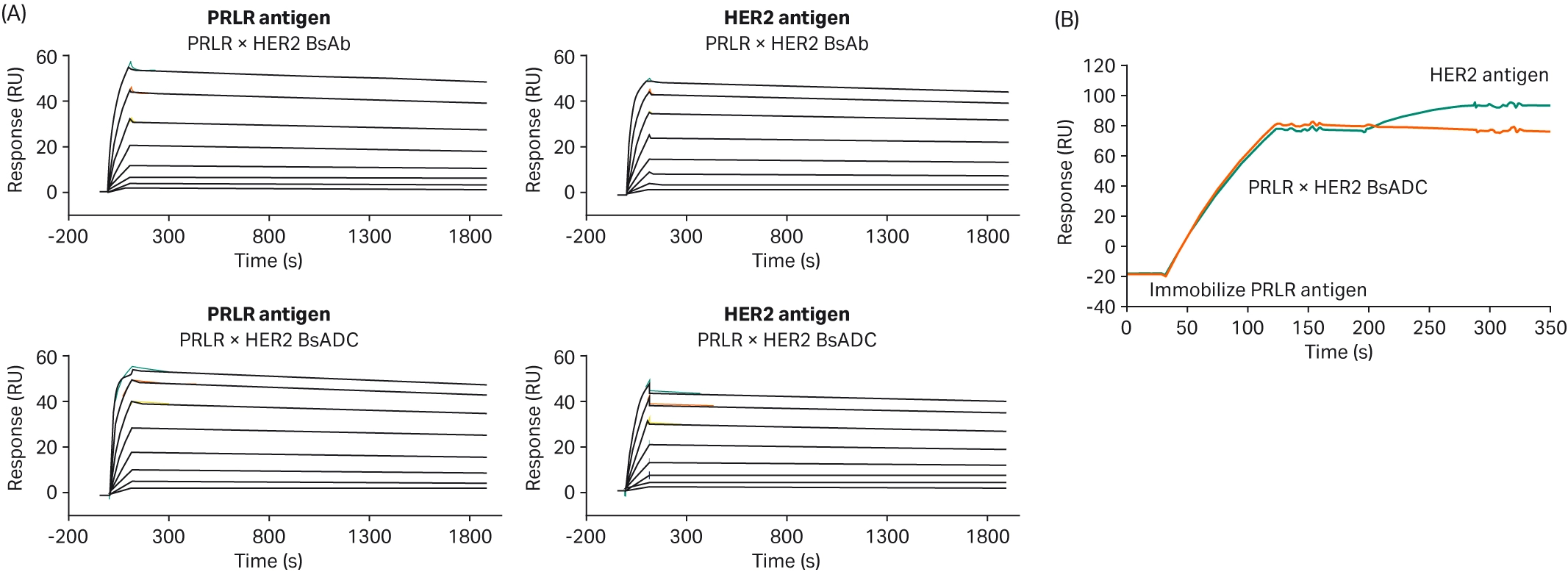
Assessing conjugation is particularly important in bispecific ADCs. Bispecific antibodies can bind to two different target proteins simultaneously, with the potential to enhance the specificity and internalization of the ADC. The additional binding event for a bispecific antibody increases the risk of the conjugation affecting the binding. Zong et al. developed a bispecific antibody targeting both PRLR and HER2, two proteins with crosstalk signaling in breast cancer (Fig 6) (15). They used Biacore™ SPR analysis to compare how the bispecific antibody binding to either the PRLR or HER2 antigens differs from the bispecific ADC binding (Fig 7A). For the PRLR antigen, they discovered that the bispecific ADC has a higher binding affinity than the bispecific antibody, whereas for the HER2 antigen, the bispecific ADC has a weaker affinity than the antibody (Table 2). Zong et al. also used SPR in a sandwich assay (Fig 7B) to confirm that the bispecific ADC could bind to both antigens simultaneously.
Fig 6. Schematic representation of the formation of the bispecific ADC.
Fig 7. SPR sensorgrams demonstrating (A) the characterization of bispecific antibody binding to either the PRLR or HER2 antigens, or (B) a sandwich assay to confirm simultaneous binding to two antigens (15).
Table 2. Binding rate constants to antigens of BsADC and the control BsAb (15)
| Antibody (antigen) | Ka (1/Ms) | Kd (1/s) | KD (nmol/L) |
|---|---|---|---|
| PRLR × HER2 BsAb (PRLR) | 2.15 × 105 | 3.17 × 104 | 1.48 |
| PRLR × HER2 BsADC (PRLR) | 6.45 × 105 | 3.47 × 104 | 0.54 |
| PRLR × HER2 BsAb (HER2) | 1.33 × 105 | 9.25 × 105 | 0.70 |
| PRLR × HER2 BsADC (PRLR) |
7.88 × 104 |
2.18 × 104 |
2.77 |
Measuring ADC binding affinity and linker stability with SPR
Linkers are required to conjugate the antibody and the cytotoxic payload. When developing ADCs, the linker structure must be carefully designed and tested to ensure that it does not impact antibody-antigen binding affinity.
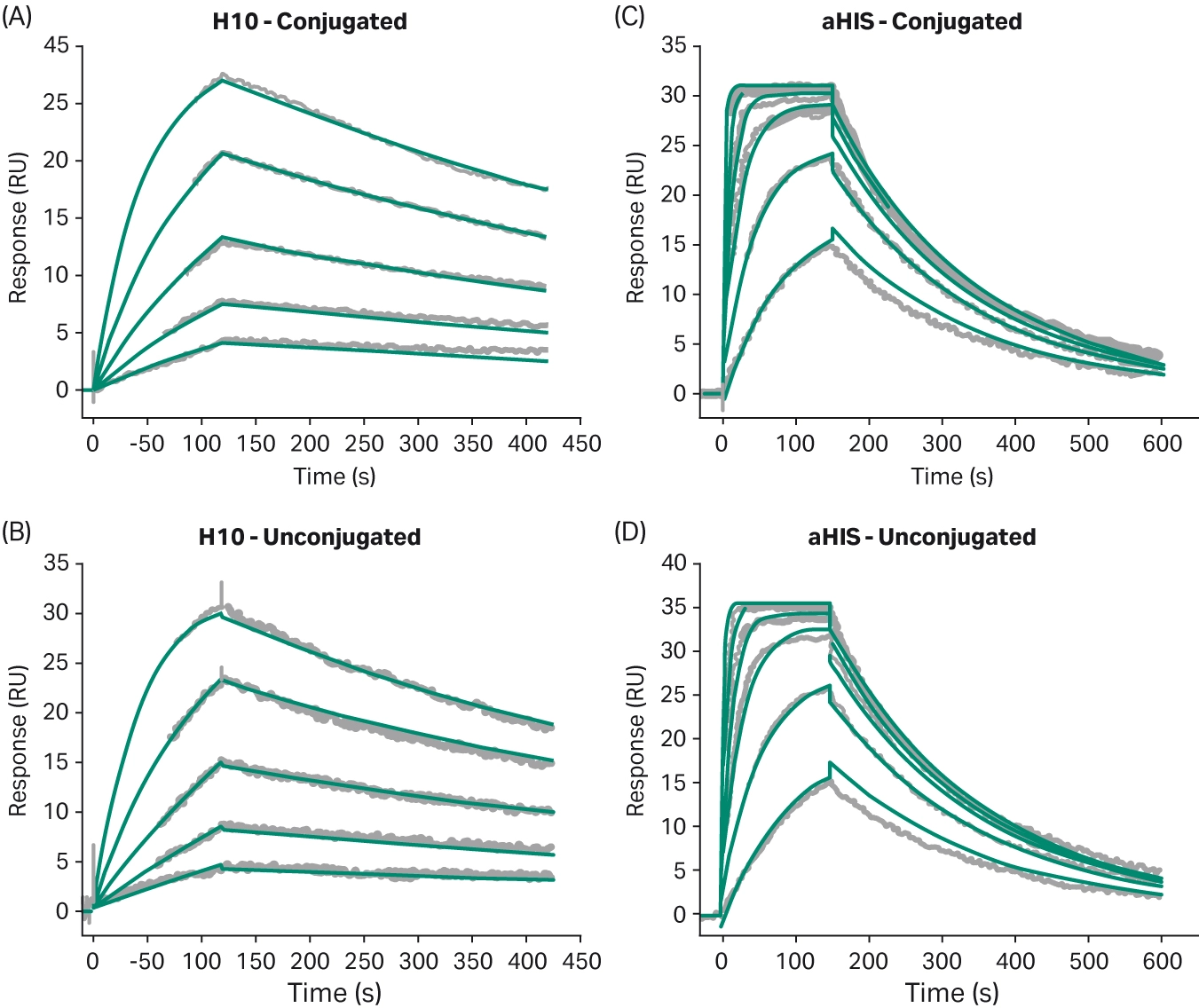
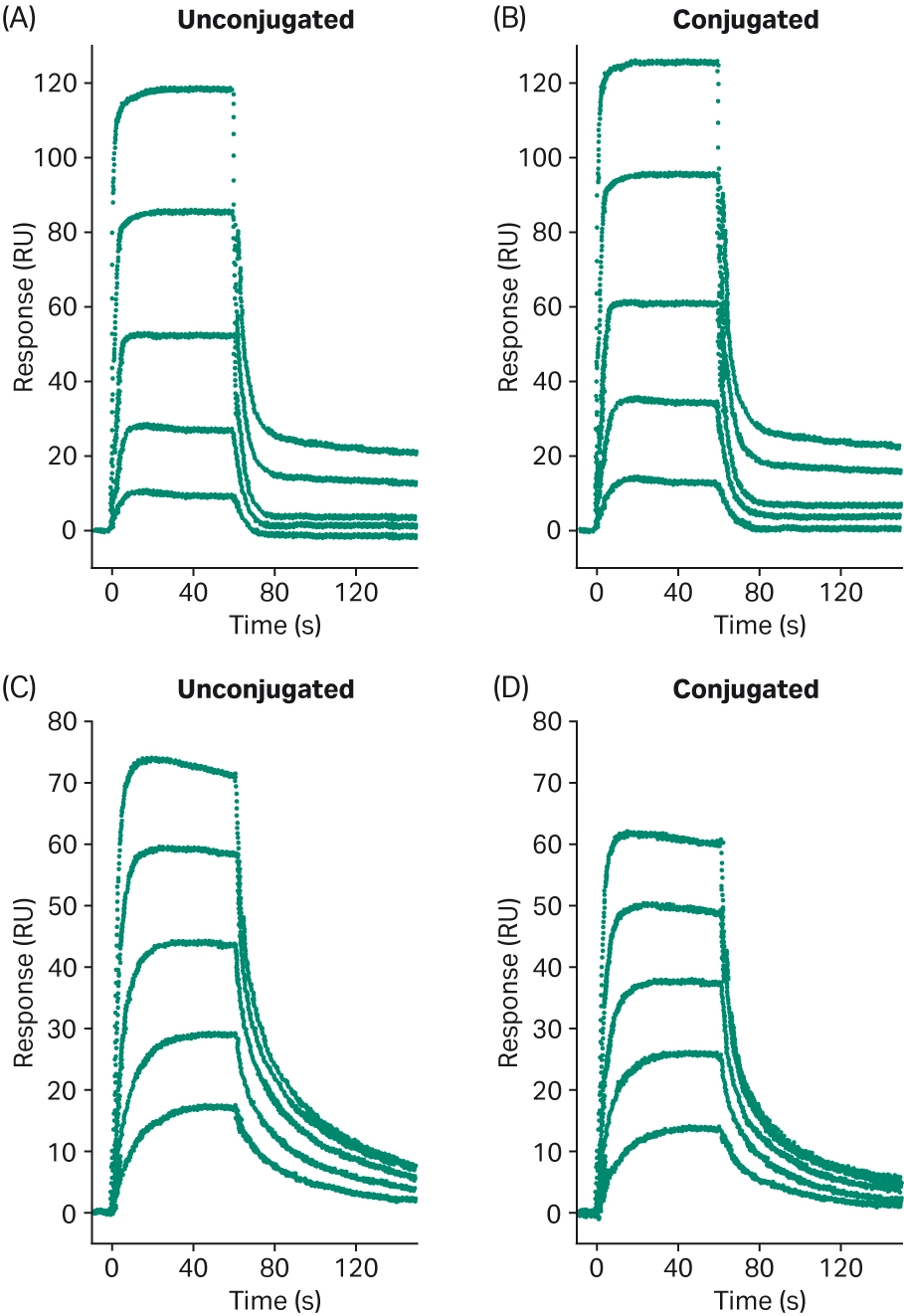
Healey et al. used Biacore™ SPR to assess whether the use of cleavable or non-cleavable linkers affected the binding kinetics of their two potential antibodies for the target antigen (4). Neither antibody showed significant differences between the cleavable and non-cleavable linkers. The effect of linker chemistry on antigen binding was also assessed by Acchione et al. using Biacore™ SPR (12). They first assessed whether linker chemistry affected antigen binding. Linkers of variable length and hydrophobicity were compared, but no significant differences were found, indicating that linker chemistry did not have an impact on antibody-antigen interactions in this experiment (Fig 8). They also assessed whether linker chemistry affected Fc receptor binding, as modifications near the Fc receptor binding site could affect binding and subsequently impact antibody function. They captured the ADC onto the immobilized antigen and then assessed the binding of the FcR to the captured ADC. While binding kinetics were mostly preserved with different linker chemistries, one of the linkers had a significant impact on FcR binding, reducing the affinity by 17-fold (Fig 9D).
Fig 8. Sensorgrams comparing binding of the ADC to the antigen using different linker chemistry (12).
Fig 9. Sensorgrams comparing ADC binding to FcR (CD32b) (12).
Assessing the effect of drug-antibody ratios on binding kinetics
The drug-to-antibody ratio (DAR) is a critical parameter in ADC development, having a direct impact on efficacy (sufficient drug transported to the cell), toxicity (higher DAR increases the risk of off-target toxicity), and pharmacokinetics/pharmacodynamics (lower DAR can result in longer systemic circulation) (16). DAR can also affect the binding kinetics of ADCs, potentially reducing the therapeutic index of the ADC.
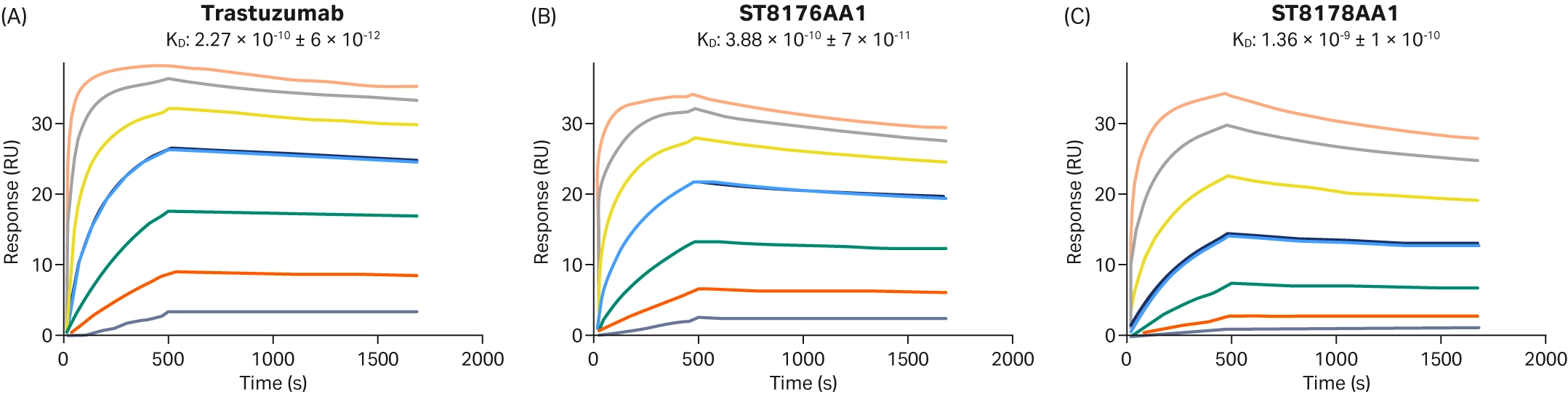
Healey et al. investigated the effect of DAR on the binding kinetics of their candidate ADC (4). They used varying molar ratios of antibody to TCEP to produce ADCs with either low or high DARs. They did not find any significant differences, suggesting that a high DAR does not impact antibody-antigen binding for this ACD. A more significant impact of DAR on binding kinetics was demonstrated by Milazzo et al (17). They investigated two new ADCs that linked the monoclonal therapeutic antibody trastuzumab to a histone deacetylase inhibitor cytotoxic payload. Each ADC had a different DAR. Biacore™ SPR was used to compare the binding kinetics of the two ADCs compared to trastuzumab binding (Fig 10). SPR results demonstrated that the ADC with lower DAR had a kinetic profile with greater similarity to trastuzumab, and from this, was chosen for further characterization.
Fig 10. Sensorgrams demonstrating that the ADC with the lower DAR (ST8176AA1) had a more similar kinetic profile than the ADC with the higher DAR (ST8178AA1) (17).
Determining how altered pH affects binding kinetics
ADCs often utilize changes in pH to control the release of the drug at the correct location (i.e., the internal cell environment). Assessing how the binding kinetics of ADCs alter at different pH is therefore crucial to ensure that the cytotoxic payload is released exclusively at the correct location.
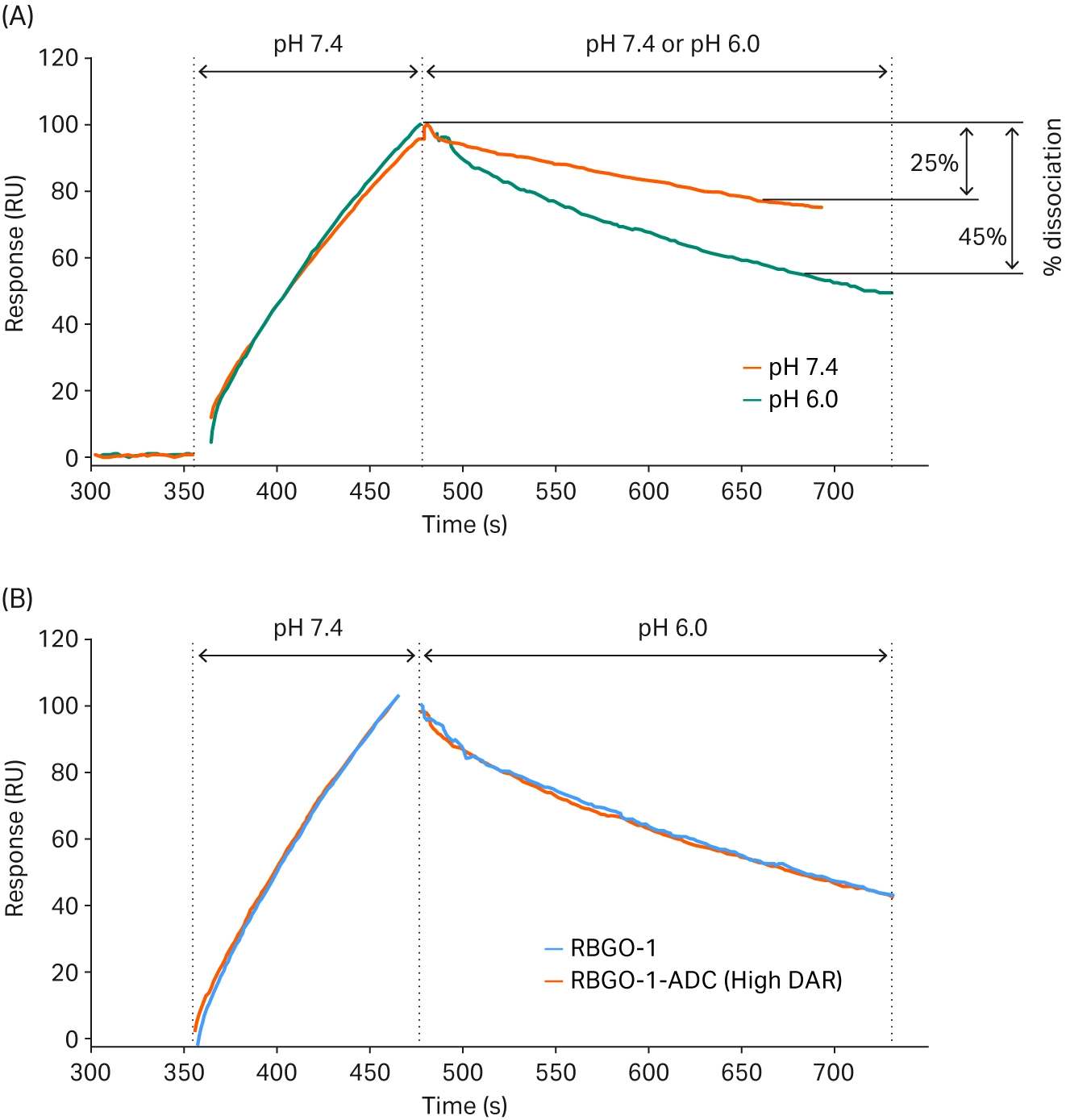
To investigate the impact of pH on the kinetics of antigen binding, Healey et al. utilized the Dual injection functionality found in certain models of Biacore™ SPR platforms (4). The use of this function enables two solutions to be injected. One solution is injected immediately after the other (with no intermediate wash steps), minimizing molecule dissociation between the two injections. In this study, the antigen was injected in a buffer at pH 7.4 to enable binding to the antibody/ADC captured onto the SPR sensor surface, immediately followed by a second injection of buffer either at pH 7.4 or pH 6 (mimicking an endosomal environment). It was found that the rate of dissociation between the antibody and antigen was increased at pH 6 compared to pH 7.4 (Fig 11). In addition, when the binding kinetics between the antibody and antigen or the ADC and antigen were compared at pH 6, no differences were observed in the rate of dissociation. Therefore, by using Biacore™ SPR, Healey et al. demonstrated that their ADC is a promising candidate with the potential to specifically target the endosomal compartment of cells as opposed to the external cell environment.
Fig 11. SPR sensorgrams showing the dissociation rate of (A) the potential antibody at different pHs, and (B) the antibody compared to the ADC at pH 6 (4).
Investigating ADC internalization estimations
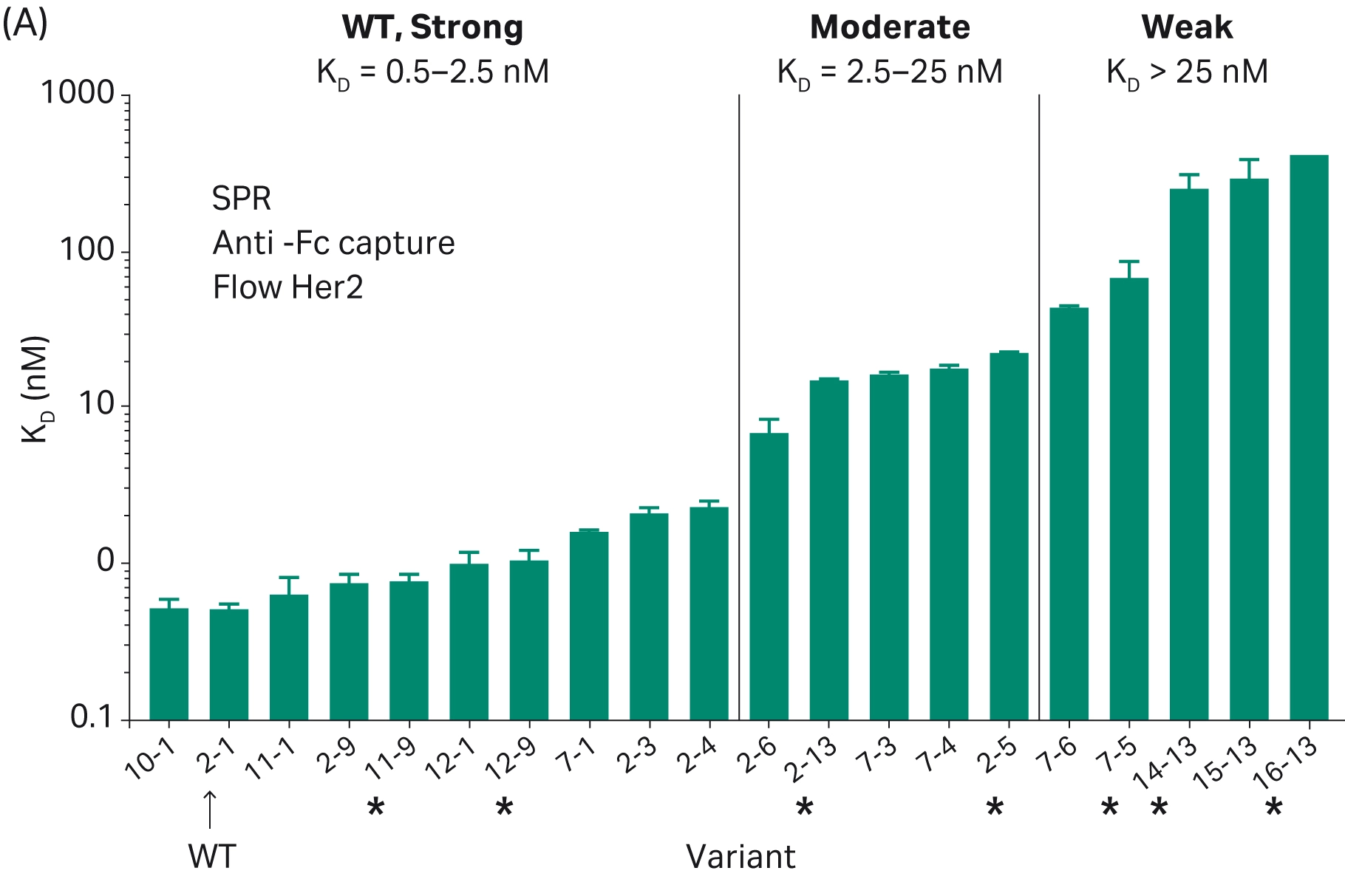
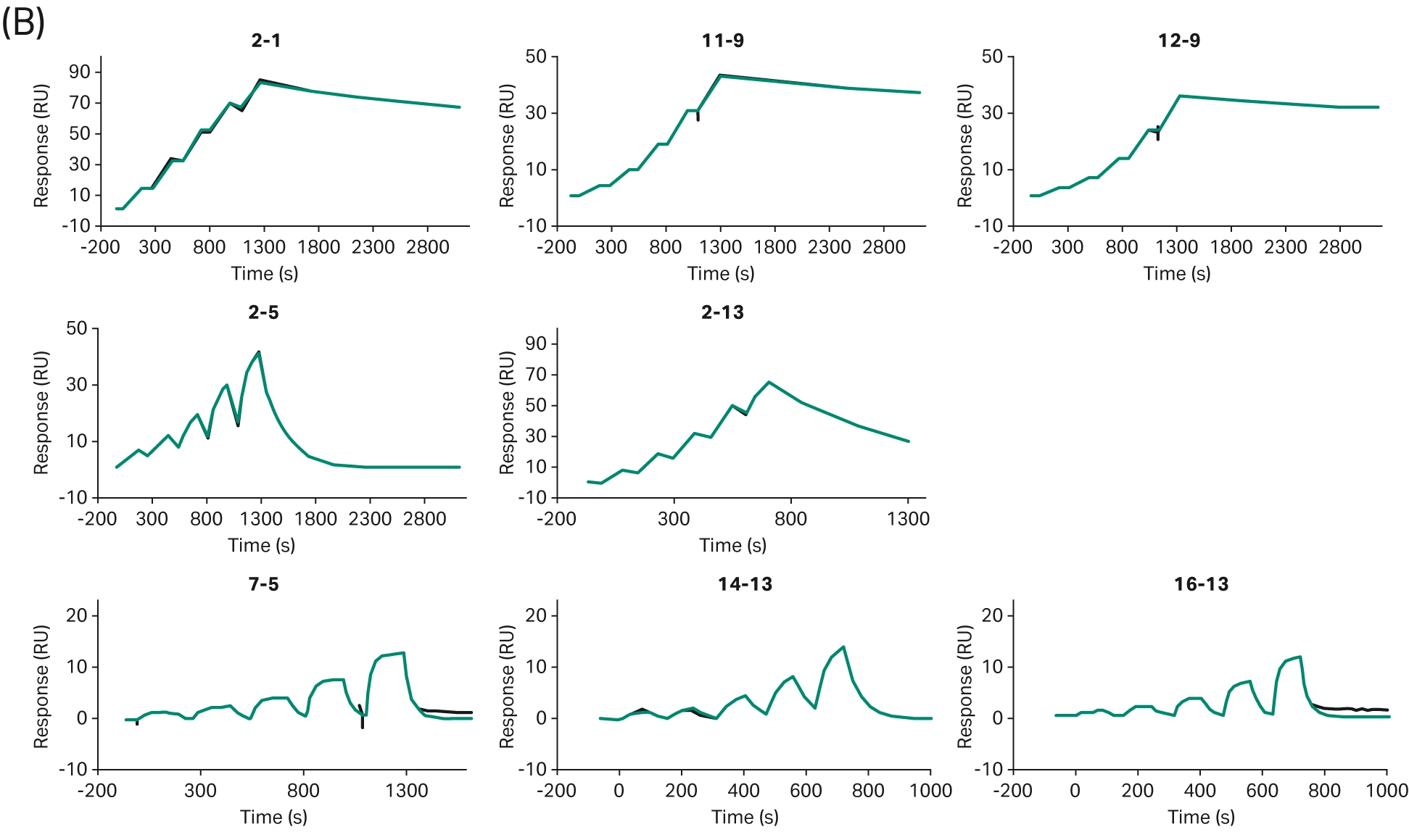
Assessing ADC internalization is an essential aspect of ADC design. Potential ADCs must be able to effectively enter cells to ensure that their cytotoxic payload is released correctly. While Biacore™ SPR technology cannot directly be used to assess internalization, binding kinetics can give an indication of internalization, where high binding affinity generally favors internalization. For example, Zwaagstra et al. used Biacore™ SPR systems to screen antibodies for ADC development, ranking the HER2 binding efficiency of a panel of antibody mutants and finding a mixture of strong, moderate, and weak binding (Fig 12) (18). Subsequent internalization analysis using high content imaging found that internalization correlated with the binding strength of the antibodies. However, the highest binding affinity does not always indicate the highest internalization. Tsumura et al. investigated how binding kinetics affected the intra-tumor distribution of ADCs. Biacore™ SPR analysis was used to assess binding kinetics, demonstrating different dissociation rates for the three ADCs against the tissue factor antigen (19). When investigated in an in vivo model, it was found that the ADC with the lowest dissociation rate (i.e., the highest binding affinity) did not show the highest anti-tumor effects (as was expected), but instead the ADC with a high dissociation rate showed significantly greater anti-tumor effects and greater distribution within the tumor.
Fig 12. (A) Ranked antibody variants based on the strength of their binding affinity. (B) Representative sensorgrams of WT/strong (top row), moderate (middle row), and weak (bottom row) (18).
Evaluating immunogenicity
Immunogenicity is caused when the patient’s body initiates an immune response to a biotherapeutic, such as an ADC, resulting in the production of anti-drug antibodies (ADAs) and potentially serious health consequences. The baseline prevalence of the production of ADAs in response to ADCs is relatively low, with one study estimating a prevalence of between 1.4% to 8.1% (20). Rigorous testing is required to accurately detect ADAs.
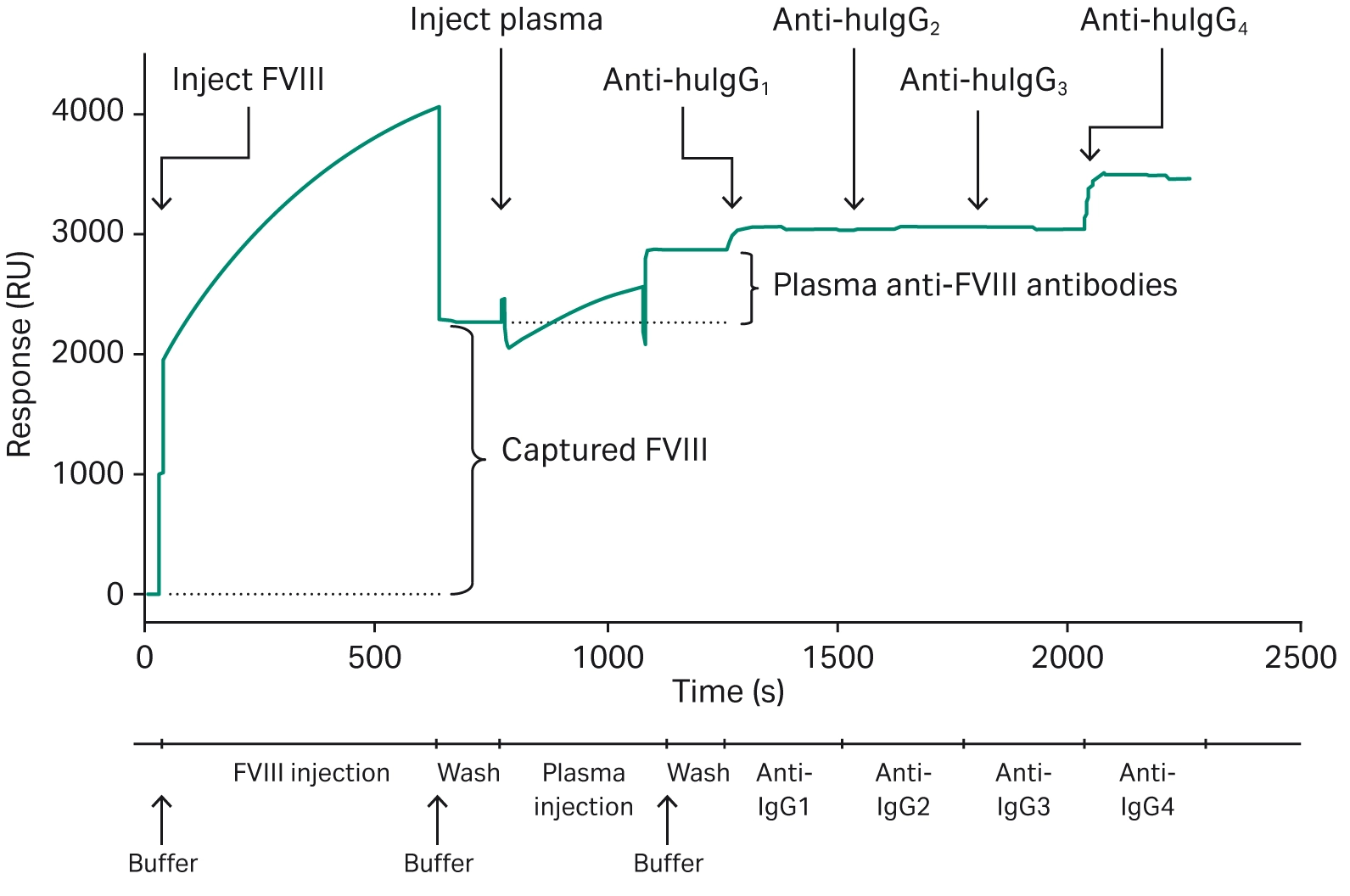
Biacore™ SPR technology can be used to screen for ADAs in patient samples, however, this has not been demonstrated specifically with ADCs. Studies investigating other drug types have shown the effectiveness of the use of Biacore™ SPR analysis compared to other ADA analyses, with an increased detection sensitivity for low-affinity ADAs and the ability to capture transient interactions with fast dissociation rates (6). Lewis et al. used a Biacore™ SPR assay to quantify all of the IgG subclasses' response in one experiment (Fig 13), demonstrating the key role that Biacore™ SPR systems can play in ADA detection and analysis (21).
Fig 13. Representative sensorgram showing the injection of the drug (FVIII), the injection of plasma and capture of ADAs, and the sequential injections of various anti-IgG antibodies (21).
The advantages of SPR techniques for antibody-drug conjugate development
Biacore™ SPR technology forms an essential tool in ADC development and characterization, providing important information on binding kinetics and the effect of conjugation, linker chemistry, pH, and DAR. Key advantages of the use of Biacore™ SPR analysis in ADC development include:
Real-time analysis
Biacore™ SPR technology assesses molecule interaction in real-time. This real-time monitoring provides quantitative data on binding kinetics as the interactions occur, enabling deeper mechanistic insights into ADC-antigen interactions, which cannot be provided by endpoint assays. In addition, real-time SPR analysis enables the detection of transient and weak interactions, such as those seen between ADAs and ADCs when assessing immunogenicity (22).
Label-free
Molecular labels, such as fluorescent tags, are often used in binding kinetic analysis. However, the binding of the label to the antigen or ADC has the potential to alter the conformation of the protein, and could alter the binding kinetics of the interaction (5). Biacore™ SPR is a label-free technique, ensuring that protein conformation and function are retained, giving accurate and reproducible results.
Expanded multiplexing capabilities
Future SPR platforms may allow simultaneous characterization of multiple binding events such as antigen interaction, Fc receptor engagement, and ADA detection within a single experiment, accelerating candidate selection.
High sensitivity for weak or transient interactions
As immunogenicity testing becomes more stringent, improvements in SPR detection limits will support early identification of transient ADA responses, mitigating clinical risk.
Versatile assay formats for complex molecule
Biacore™ SPR technology accommodates bispecific ADCs and multi-component systems, enabling researchers to study simultaneous binding events and confirm functional integrity after conjugation.
High-throughput
Modern Biacore™ SPR systems facilitate high-throughput experimentation, enabling rapid drug screening (23).
AI incorporation
Biacore™ SPR produces large datasets, which often require sufficient time and experience to analyze. However, advancements in artificial intelligence (AI) and machine learning have enabled incorporation into Biacore™ software, and can be used to optimize and enhance data interpretation. Biacore Intelligent Analysis™ is an optional machine learning software extension that can assist researchers by improving flexibility, streamlining workflows, and saving up to 80% of data evaluation time (24).
Integration with other digital lab tools
The integration of Biacore™ SPR systems with digital lab tools, such as laboratory information management systems and electronic lab notebooks, is enabled by Biacore™ Insight Software. The integrated application programming interface (API) enables the automated integration of the Biacore™ SPR system data directly into lab information management systems. This software also supports customizable data export in standard formats to allow connectivity throughout digital lab infrastructures.
Conclusion
Biacore™ SPR systems form an essential tool to develop and validate ADCs. They enable the characterization of binding kinetics, including how conjugation, linker chemistry, pH, and DAR can affect the interaction between the ADC and the target protein, giving further indication of drug efficacy and potency. Multiple advantages are provided through the use of Biacore™ SPR to research ADCs, including real-time, label-free analysis with sufficient sensitivity to detect low-affinity, transient reactions. Recent advances in Biacore™ SPR technology have also enabled the incorporation of AI and systems capable of integration with other digital tools.
References
- Fu Z, Li S, Han S, Shi C, Zhang Y. Antibody drug conjugate: the “biological missile” for targeted cancer therapy. Sig Transduct Target Ther. 2022;7(1):93. doi:10.1038/s41392-022-00947-7
- Pettinato MC. Introduction to Antibody-Drug Conjugates. Antibodies (Basel). 2021;10(4):42. doi:10.3390/antib10040042
- PEG P. Antibody–Drug Conjugates Approved by FDA Through 2025. Precise PEG. Accessed September 5, 2025. https://precisepeg.com/blogs/posts/antibody-drug-conjugates-approved-through-2025
- Healey GD, Frostell A, Fagge T, Gonzalez D, Conlan RS. A RAGE-Targeted Antibody-Drug Conjugate: Surface Plasmon Resonance as a Platform for Accelerating Effective ADC Design and Development. Antibodies (Basel). 2019;8(1):7. doi:10.3390/antib8010007
- Bobone S, Storti C, Fulci C, et al. Fluorescent Labeling Can Significantly Perturb Measured Binding Affinity and Selectivity of Peptide-Protein Interactions. J Phys Chem Lett. 2024;15(40):10252-10257. doi:10.1021/acs.jpclett.4c01767
- Beeg M, Burti C, Allocati E, et al. Surface plasmon resonance unveils important pitfalls of enzyme-linked immunoassay for the detection of anti-infliximab antibodies in patients’ sera. Sci Rep. 2021;11(1):14976. doi:10.1038/s41598-021-94431-x
- Barbosa MDFS, Gokemeijer J, Martin AD, Bush A. Altering drug tolerance of surface plasmon resonance assays for the detection of anti-drug antibodies. Analytical Biochemistry. 2013;441(2):174-179. doi:10.1016/j.ab.2013.07.019
- Lin S, Zhang Y, Yao J, et al. DB-1314, a novel DLL3-targeting ADC with DNA topoisomerase I inhibitor, exhibits promising safety profile and therapeutic efficacy in preclinical small cell lung cancer models. J Transl Med. 2024;22:766. doi:10.1186/s12967-024-05568-y
- Li X, Zhou S, Abrahams CL, et al. Discovery of STRO-002, a Novel Homogeneous ADC Targeting Folate Receptor Alpha, for the Treatment of Ovarian and Endometrial Cancers. Mol Cancer Ther. 2023;22(2):155-167. doi:10.1158/1535-7163.MCT-22-0322
- Junutula JR, Raab H, Clark S, et al. Site-specific conjugation of a cytotoxic drug to an antibody improves the therapeutic index. Nat Biotechnol. 2008;26(8):925-932. doi:10.1038/nbt.1480
- Onyido EK, James D, Garcia-Parra J, et al. Elucidating Novel Targets for Ovarian Cancer Antibody–Drug Conjugate Development: Integrating In Silico Prediction and Surface Plasmon Resonance to Identify Targets with Enhanced Antibody Internalization Capacity. Antibodies (Basel). 2023;12(4):65. doi:10.3390/antib12040065
- Acchione M, Kwon H, Jochheim CM, Atkins WM. Impact of linker and conjugation chemistry on antigen binding, Fc receptor binding and thermal stability of model antibody-drug conjugates. MAbs. 2012;4(3):362-372. doi:10.4161/mabs.19449
- Cardillo TM, Govindan SV, Sharkey RM, et al. Sacituzumab Govitecan (IMMU-132), an Anti-Trop-2/SN-38 Antibody-Drug Conjugate: Characterization and Efficacy in Pancreatic, Gastric, and Other Cancers. Bioconjug Chem. 2015;26(5):919-931. doi:10.1021/acs.bioconjchem.5b00223
- Chen L, Wang L, Shion H, et al. In-depth structural characterization of Kadcyla® (ado-trastuzumab emtansine) and its biosimilar candidate. MAbs. 2016;8(7):1210-1223. doi:10.1080/19420862.2016.1204502
- Zong HF, Zhang BH, Zhu JW. Generating a Bispecific Antibody Drug Conjugate Targeting PRLR and HER2 with Improving the Internalization. Pharmaceutical Fronts. 2022;04:e113-e120. doi:10.1055/s-0042-1749334
- Mahmood I. Clinical Pharmacology of Antibody-Drug Conjugates. Antibodies (Basel). 2021;10(2):20. doi:10.3390/antib10020020
- Milazzo FM, Vesci L, Anastasi AM, et al. ErbB2 Targeted Epigenetic Modulation: Anti-tumor Efficacy of the ADC Trastuzumab-HDACi ST8176AA1. Front Oncol. 2020;9:1534. doi:10.3389/fonc.2019.01534
- Zwaagstra JC, Sulea T, Baardsnes J, et al. Binding and functional profiling of antibody mutants guides selection of optimal candidates as antibody drug conjugates. PLoS One. 2019;14(12):e0226593. doi:10.1371/journal.pone.0226593
- Tsumura R, Manabe S, Takashima H, Koga Y, Yasunaga M, Matsumura Y. Influence of the dissociation rate constant on the intra-tumor distribution of antibody-drug conjugate against tissue factor. Journal of Controlled Release. 2018;284:49-56. doi:10.1016/j.jconrel.2018.06.016
- Carrasco-Triguero M, Dere RC, Milojic-Blair M, et al. Immunogenicity of Antibody–drug Conjugates: Observations Across 8 Molecules in 11 Clinical Trials. Bioanalysis. 2019;11(17):1555-1568. doi:10.4155/bio-2018-0259
- Lewis KB, Hughes RJ, Epstein MS, et al. Phenotypes of Allo- and Autoimmune Antibody Responses to FVIII Characterized by Surface Plasmon Resonance. PLoS One. 2013;8(5):e61120. doi:10.1371/journal.pone.0061120
- Livnat Levanon N, Vigonsky E, Lewinson O. Real time measurements of membrane protein:receptor interactions using Surface Plasmon Resonance (SPR). J Vis Exp. 2014;(93):e51937. doi:10.3791/51937
- Schneider CS, Bhargav AG, Perez JG, et al. Surface plasmon resonance as a high-throughput method to evaluate specific and non-specific binding of nanotherapeutics. J Control Release. 2015;219:331-344. doi:10.1016/j.jconrel.2015.09.048
- Machine learning drives smarter SPR analysis. Accessed September 8, 2025. https://www.selectscience.net/article/machine-learning-drives-smarter-spr-analysis
Images (except Fig 2) have been adapted from the respective original author's works listed in the references under Creative Commons License 4.0.
CY57750