This whitepaper is part of our Biacore™ SPR applications series.
In this series, we explore how Biacore™ SPR technology supports a wide range of applications—from membrane protein analysis to fragment screening, ADCs, and beyond. Each article highlights real-world data and use cases to help researchers unlock the full potential of SPR.
Key takeaways:
- Small molecules are increasingly being used in drug discovery, making up 90% of marketed drugs.
- Multiple challenges in analyzing small-molecule binding interactions create a hurdle to small-molecule drug discovery.
- Biacore™ SPR technology overcomes analytical hurdles through its sensitivity, versatility, and compatibility with challenging targets. These aspects enable its use in fragment-based drug design, drug screening, kinetic characterization by providing mechanistic insights and binding site mapping.
Introduction to small molecules
Small molecules are synthetic or natural low molecular weight (LMW) compounds that interact with specific biological targets. Their small size allows them to move through the body easily, including through the bloodstream to their target site and through cell membranes to access intracellular targets (1). Small molecules can produce a therapeutic response within the body using several mechanisms, including inhibiting enzymes, acting as receptor agonists or antagonists, or modulating ion channels. These qualities have led to small molecules playing a pivotal role in drug discovery, currently making up 90% of marketed drugs. As such, the aim of pre-clinical work is to identify and characterize the interactions of small molecules with their target protein (2). This process involves:
- Screening libraries of small molecules or small molecule fragments to identify potential candidates
- Validating the engagement of the candidate molecules with the target proteins
- Identifying the molecular mode of action
- Obtaining a detailed characterization of compound binding kinetics
Analytical hurdles to drug discovery using small molecules
Complex chemical space
Small molecule drug discovery involves screening libraries containing millions of small molecules, which is a small proportion of the total possible number of small drug-like molecules (3). To perform a more thorough search of potential drug candidates, fragment-based drug discovery (FBDD) has been introduced. The total possible number of fragments is significantly less than for small molecules, resulting in efficient sampling of the chemical space. Therefore, FBDD generally screens significantly fewer molecules compared to small-molecule screening.
Challenging kinetics and complex drug targets
Small molecules have unique properties that complicate the characterization of the kinetics of target protein binding, which is a crucial aspect of drug screening and discovery. Small molecules and fragments often exhibit weak binding affinity and are difficult to detect using traditional screening mechanisms (4). In addition, the kinetics of small molecule reactions are often rapid and transient, making standard endpoint assays insufficient to capture accurate data. Small molecules also have a low molecular weight, which can result in low signal-to-noise ratios and difficulty identifying candidates for further drug development. Furthermore, the use of labels is a common aspect of many techniques used to analyze binding kinetics, which can potentially interfere with the interaction and exacerbate concerns that the label may alter binding kinetics or protein conformation.
Many emerging drug targets are considered ‘undruggable’ due to their lack of accessible binding pockets or belonging to homologous families where binding specificity is a challenge (5). These issues make it difficult to screen small molecules for potential binding candidates. Innovative design approaches may be needed to identify potential targets.
Surface plasmon resonance technology overview
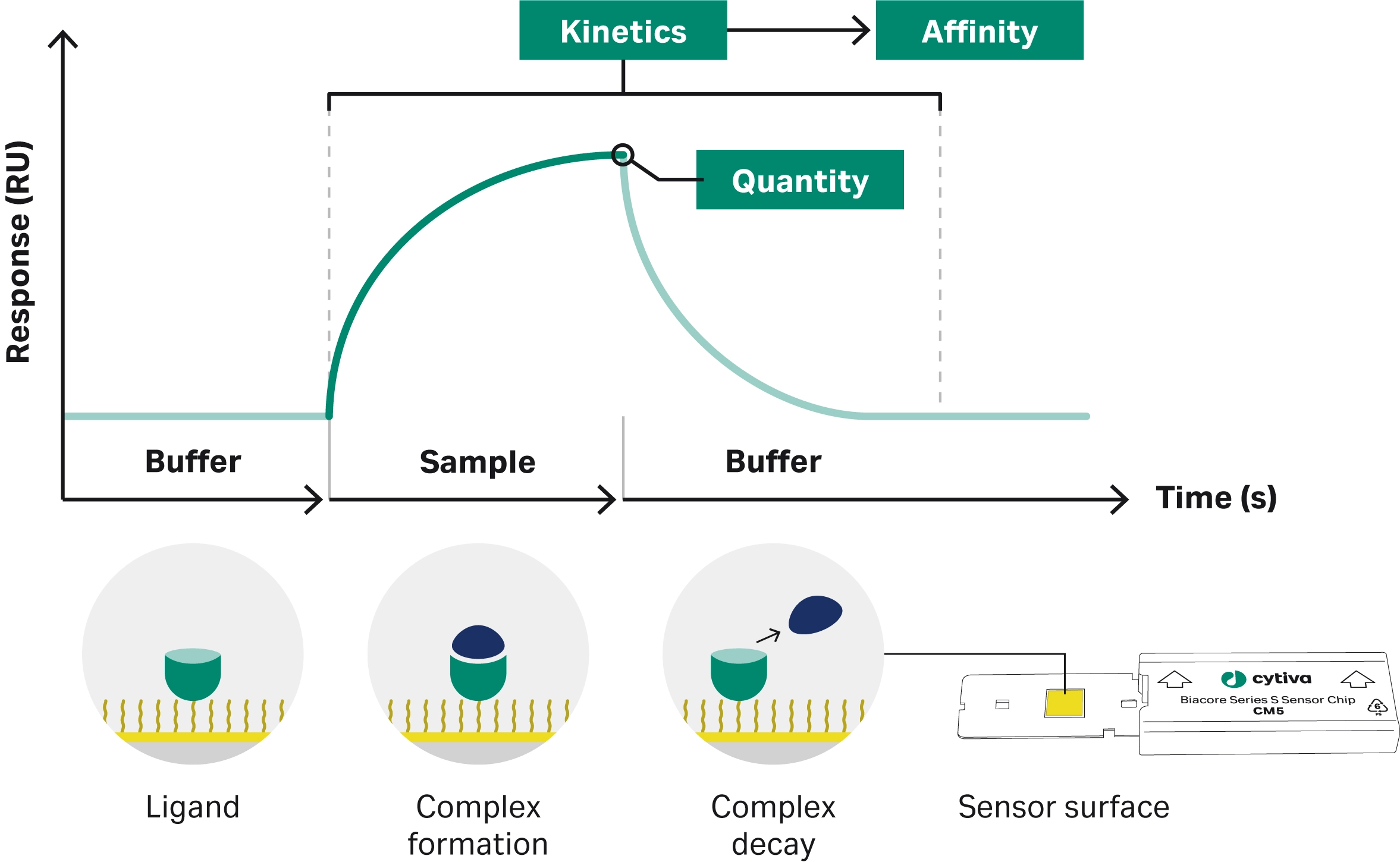
Surface plasmon resonance (SPR) is a technique used to analyze biomolecular interactions. Plasmons are electron density waves generated when polarized light hits an SPR sensor chip at an angle of total internal reflection. At a specific resonance angle, energy is absorbed by the plasmons. When molecules bind to the target on the sensor chip, the mass changes and causes a shift in the refractive index and resonance angle (6, 7). The SPR angle change is proportional to mass change on the sensor surface. The SPR angle changes are converted to binding responses that are presented in sensorgrams (Fig 1).
Target molecules are attached to the SPR sensor chip surface. The potential binding partner is injected over the immobilized target. As the sample binds to the target molecule, the change in mass at the surface causes a change to the resonance angle. This is recorded in real time throughout the experiment to produce a sensorgram which can then be analyzed to determine association and dissociation rates, binding affinity, specificity, and concentration (6).
Fig 1. An example sensorgram. The bars below the graph represent the solutions passing over the sensor surface.
Biacore™ SPR systems offer unmatched sensitivity, label-free detection, and real-time kinetic analysis. Designed for flexibility and precision, Biacore SPR technology enables researchers to confidently characterize molecular interactions—even with low-affinity, fast-dissociating compounds or challenging targets. Whether screening fragments, mapping binding sites, or optimizing drug candidates, Biacore SPR technology delivers the throughput, capacity, and high-quality data you need to accelerate LMW drug discovery.
Biacore™ SPR system features
Biacore SPR systems are industry-leading platforms for label-free interaction analysis using SPR technology. Designed to deliver high-quality, information-rich data, Biacore SPR system answers critical questions about molecular binding—accelerating your research toward meaningful conclusions.
With advanced features for high-throughput and high-sensitivity analysis, Biacore SPR system enables real-time evaluation of affinity, kinetics, epitope binning, and concentration across a broad molecular weight range using minimal sample volumes.
Biacore™ Insight Software provides unified instrument control and data evaluation with optional application-specific extensions. It includes machine learning-based tools and seamless integration capabilities for automated workflows and cross-platform data management to enhance efficiency and reproducibility in your lab.
The use of Biacore SPR system with low-molecular-weight chemical compounds
SPR is an effective tool to assess the binding kinetics of interactions between low molecular-weight compounds and target proteins.
How to use SPR for small molecule drug screening?
Fragment screening has increased in popularity as an effective screening method covering a wider chemical space than other screening methods.
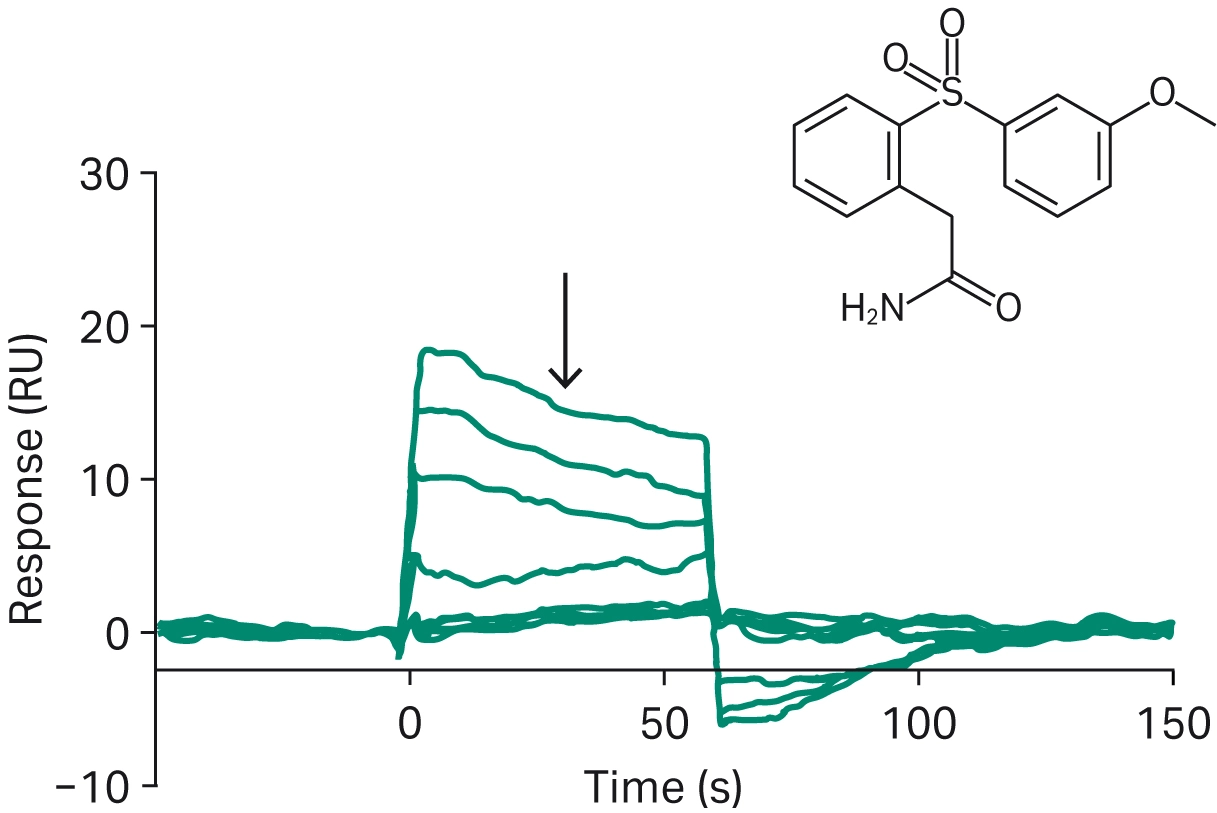
Biacore SPR system is a favored tool for screening fragment libraries and identifying possible hits for further research. Huber et al. used Biacore SPR system to screen a fragment library of 6369 compounds for binding to neurotensin 1, a drug target for various central nervous system diseases (8). They identified 44 hits, which were further narrowed down to four hits after follow-up experiments using nuclear magnetic resonance (NMR) (Fig 2). Heinrich et al. also screened a fragment library using Biacore SPR system, identifying bicyclic scaffolds binding to the hinge region of focal adhesion kinase (9). Li et al. used a Biacore SPR fragment screen to identify two fragments that bind to a binding pocket of the catalytic subunit of ricin, with implications for inhibiting poisoning by the ricin holotoxin (10). Luttens et al. used a novel virtual screening method to perform a primary screen of 235 million compounds for binding to the SARS-CoV-2 main protease (11). Biacore SPR system was then used in a secondary screening of 100 compounds selected by the primary screening to identify nineteen compounds showing measurable binding to the target protease. Biacore SPR technology is considered one of the best methods for fragment-based drug discovery.
Fig 2. Representative SPR sensorgram for fragment 2 (8).
Addressing complex drug targets with SPR
Challenging drug targets complicate fragment screens and can result in the designation of ‘undruggable’ targets. FitzGerald et al. demonstrated the use of Biacore SPR technology and multiplexed strategies to screen fragment libraries against five challenging targets (12). They include:
- Large and structurally dynamic targets
- Targets found within multi-protein complexes
- Unstable targets
- Targets containing intrinsically disordered regions
- Targets prone to aggregation
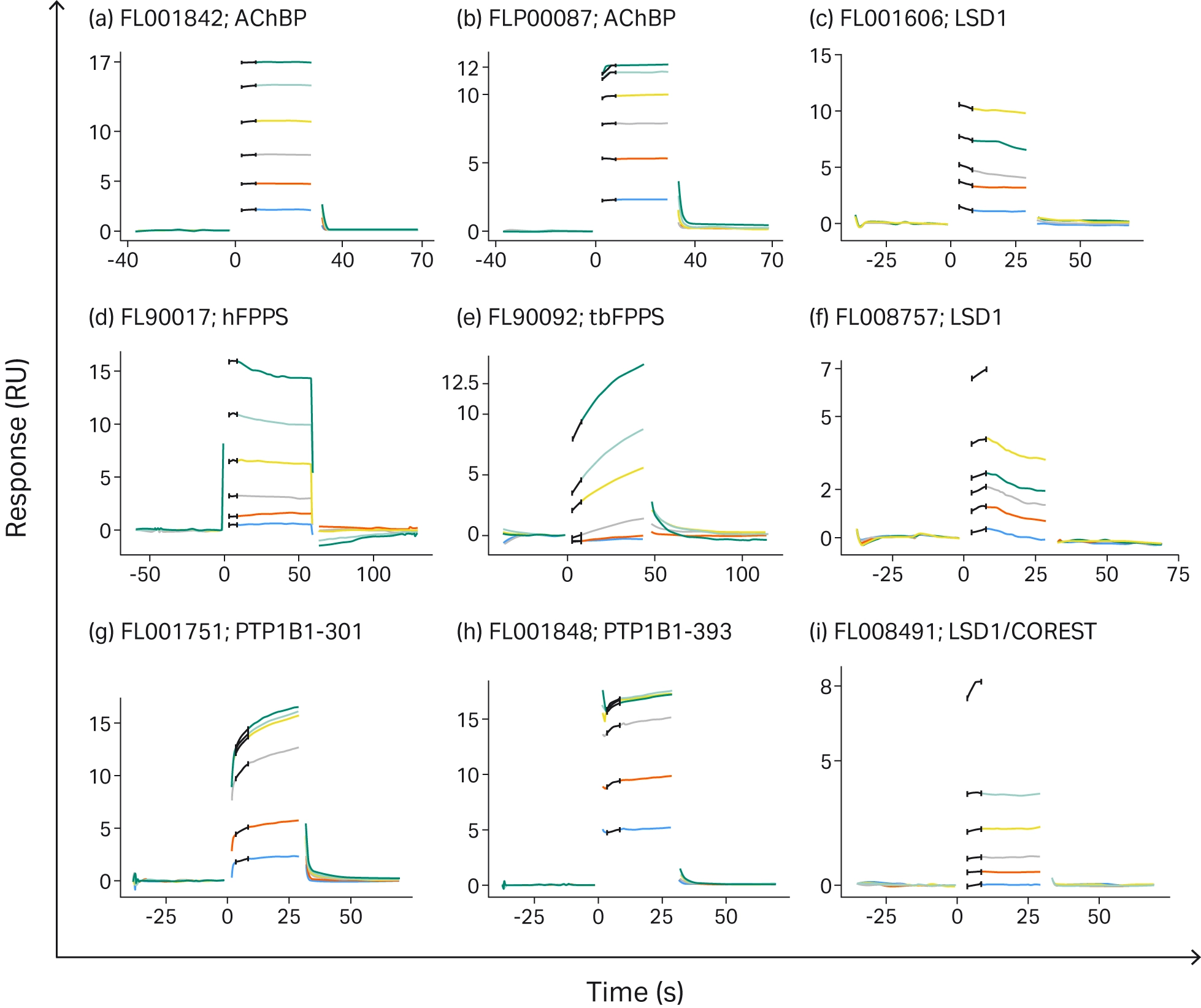
For each of the targets, fragment hits were successfully confirmed, and binding affinity data were also determined for many of the targets (Fig 3).
Fig 3. Sensorgrams of various SPR fragment screens against challenging targets. Sensorgrams a-c are typical fragment sensorgrams, with fast association and dissociation kinetics. Sensorgrams d-i demonstrate fragments showing non-ideal interaction characteristics (12).
Kinetic and affinity characterization of small molecules with SPR
Small-molecule drugs function through binding to their target protein, causing effects such as enzyme inhibition, receptor antagonism, or ion channel modulation. Therefore, when developing small-molecule drug candidates, it is essential to establish reliable target protein-small molecule binding assays to determine the kinetics of protein binding and the binding affinity.
Li et al. used Biacore SPR system to assess their efforts to optimize the affinity and inhibitory activity of their hit identified in a fragment screen (10). The researchers used a structure-based design to obtain four new compounds based on the initial hit. Biacore SPR system was then used to assess the binding of the four compounds to the target protein, with each showing a higher affinity than the original compound. Heinrich et al. also carried out chemical modifications of their prime candidate identified in a fragment screen, before using Biacore SPR system to assess how the modifications affected binding kinetics to their target protein (9). Kim et al. used Biacore SPR system to assess the binding kinetics of a series of small-molecule glycomimetics to galectin-3 (13). Confirming target engagement is a crucial step in developing small molecules for use as drugs.
Target residence time and mechanistic insights
SPR can be used to provide mechanistic insights into small-molecule binding interactions, including residence time and predicting cell potency. Specialized methods to assess assays with binding kinetics outside of the ranges measured in standard SPR experiments can be employed.
In a study investigating the potential of threonine tyrosine kinase (TTK) inhibitors to treat aggressive cancers, Uitdehaag et al. used Biacore SPR system to assess small-molecule compound binding to TTK (14). Twelve TTK small-molecule inhibitors were analyzed, cellular assays were carried out to assess the cell potency (IC50), and binding kinetics were calculated. The binding kinetics were correlated to cellular potency to determine which parameter had the greatest effect on inhibition. They found that while the affinity showed good correlation with cell potency and could broadly predict the cell potency, the residence time (koff) had a significantly higher correlation with cell potency. The researchers used these insights to develop TTK inhibitors with longer residence times. This study demonstrated the importance of determining kinetic parameters for use as optimization metrics in drug development.
Thinn et al. used a specialized SPR method to assess bimolecular interactions with very slow dissociation rate constants (15). In preliminary studies, the researchers found that their prospective small molecules created very strong binding complexes with the target proteins. However, due to their binding stability, they had dissociation rates slower than the limit of resolution. This prevented the further discovery of lead candidates using target-engagement experiments. In this study, they used an SPR chaser assay to investigate the dissociation rate of small molecules that tightly bind to the target protein. In a chaser assay, a competitive molecule, known as the chaser, is injected at specific time intervals during the dissociation phase of SPR. It binds to free-binding sites on the target protein (which are available due to dissociation of the small molecules) and enable determination of fractional occupancy, which reflects the dissociation of the small molecules from the target protein (16). Thinn et al. used the chaser assay to determine the dissociation rate of small molecules tightly bound to their target proteins.
Selectivity and binding site mapping
Biacore SPR systems can assess small-molecule binding selectivity and map the binding site location using specialized methods.
FitzGerald et al. used Biacore SPR systems to assess the binding sites on a protein target found within a protein complex (12). They performed a fragment screen against both the target protein alone and against the target protein within its protein complex. A larger number of hits were confirmed against the target protein alone than when in the complex, suggesting that the interface between the two proteins contains fragment binding sites. FitzGerald et al. also investigated how SPR can be used to determine fragment hits specific to different protein domains. They used two engineered forms of the target protein, one containing the catalytic domain and one additionally containing an intrinsically disordered region. Groups of hits unique to the catalytic domain, unique to the intrinsically disordered region, and common to both domains were identified.
Biacore SPR systems can be used to map binding sites by using a technique called biophysical mapping. Zhukov et al. demonstrated the use of this technique to map binding sites to a stabilized receptor (StaR; GPCR receptors minimally engineered for thermostability external to the membrane) of the A2A receptor, which is a promising drug target in Parkinson’s disease (17). They introduced single mutations at suggested small-molecule binding sites and immobilized the panel of mutants onto SPR sensor chips. Biacore SPR technology was then used to determine the kinetics of the binding of small molecules to each mutant. The SPR data were combined with molecular modeling and docking to map small-molecule binding sites and produce a 3-D map of ligand binding.
The advantages of SPR in studying low-molecular-weight chemical compounds
Small molecules currently serve as the main avenue for drug discovery. However, they have unique properties that complicate binding kinetics assay analysis including weak binding affinities, rapid and transient kinetics, low signal-to-noise ratio, and necessity for label-free analysis. Biacore SPR sytem can be used for multiple aspects of small protein research including SPR fragment screening, fragment-based drug design, quantifying target binding, and mapping binding sites. Key advantages of the use of Biacore SPR system in small molecule research include:
High sensitivity
Fragments and drug compounds are intrinsically small in mass, which demands exquisite signal-to-noise ratios. Small molecule interactions often have weak affinity, making binding kinetics difficult to assess using traditional techniques. Biacore SPR system is highly sensitive, with the ability to reliably detect interactions and measure binding kinetics regardless of molecular weight.
Label-free
Molecular labels are often used in binding analysis, but binding of the label to the antigen or target protein can alter protein conformation and give inaccurate binding kinetics data (18). This is a particular concern with the use of small molecules, as the size ratio between the molecular tags and the small molecule increases the risk of altered binding (4). SPR is a label-free technique, so small molecules and their target proteins retain their native function and conformation for accurate and reproducible results.
Real-time analysis
Small molecules often have rapid and transient kinetics that can be missed if endpoint assays are being used (19). Real-time analysis of low molecular weight compounds can be performed using Biacore SPR system. The assessment of molecular interactions in real-time provides rapid and transient interaction data that is collected as the interaction occurs, so you get accurate characterization of small-molecule binding kinetics.
High-throughput
High-throughput techniques are essential for screening as they enable large fragment libraries to be assessed efficiently. Modern Biacore SPR system facilitates high-throughput experimentation for rapid screening and hit identification (20).
AI incorporation
The large datasets produced by Biacore SPR analysis can be challenging and time-consuming to analyze, particularly for inexperienced users. However, the incorporation of artificial intelligence (AI) and machine learning into Biacore Insight software helps users optimize and enhance data interpretation (21). Biacore Intelligent Analysis™ software is an optional supervised machine learning software extension capable of streamlining data analysis workflows by saving 80% of the data analysis time, and providing faster, unbiased identification of true hits. In addition to the integration of AI, Biacore SPR technology is increasingly being integrated into AI-driven workflows for small molecule drug discovery for powerful capabilities in both automation and data analysis.
Compatibility with challenging targets
Biacore SPR system is versatile, providing experimental flexibility and enabling researchers to investigate a range of mechanistic questions using a range of challenging targets. On the optimization of immobilization conditions, Biacore SPR system can be used with many protein targets including disordered, aggregation-prone, unstable, and membrane proteins (12, 17, 22).
Integration with other digital lab tools
Biacore SPR systems can be integrated with other digital lab tools, including electronic lab notebooks and laboratory information management systems. This is straightforward to perform using Biacore Insight data integration extension, which supports customizable data export in standard formats to allow inter-lab connectivity.
Conclusion
Small molecules present unique challenges in drug discovery, from weak binding affinities to complex target environments. Biacore SPR system addresses these hurdles with sensitivity, high-throughput, and label-free analysis assay tools. The associated consumables, software, and applications expertise assist in efficient and reliable fragment or small molecule screening, hit identification, target binding quantification, mechanistic insights, and binding site mapping. Biacore SPR real-time analysis measures rapid and transient interactions and provides a wealth of data to guide drug development. With advanced features like AI-driven data interpretation and compatibility with digital lab tools, Biacore SPR system is not just a biosensor; it’s a strategic tool for small molecule innovation.
Contact Cytiva today to learn more about how Biacore SPR systems can help you develop and validate small molecule screening and drug characterization assays to accelerate drug discovery.
FAQ
What is SPR?Surface plasmon resonance (SPR) is a label-free technique used to study biomolecular interactions in real time. It detects changes in mass near a sensor surface when molecules bind, providing insights into binding kinetics, affinity measurements, and concentration. SPR is widely used in drug screening and discovery and protein characterization due to its sensitivity and versatility.
References
- Southey M, Brunavs M. Introduction to small molecule drug discovery and preclinical development. Frontiers in drug discovery. 2023;3. doi:10.3389/fddsv.2023.1314077
- Kaminski T, Gunnarsson A, Geschwindner S. Harnessing the Versatility of Optical Biosensors for Target-Based Small-Molecule Drug Discovery. ACS Sens. 2017;2(1):10-15. doi:10.1021/acssensors.6b00735
- Erlanson DA, Fesik SW, Hubbard RE, Jahnke W, Jhoti H. Twenty years on: the impact of fragments on drug discovery. Nat Rev Drug Discov. 2016;15(9):605-619. doi:10.1038/nrd.2016.109
- Chieng A, Wan Z, Wang S. Recent Advances in Real-Time Label-Free Detection of Small Molecules. Biosensors. 2024;14(2):80. doi:10.3390/bios14020080
- Xie X, Yu T, Li X, et al. Recent advances in targeting the “undruggable” proteins: from drug discovery to clinical trials. Signal Transduct Target Ther. 2023;8(1):335. doi:10.1038/s41392-023-01589-z
- Article: Surface plasmon resonance. Cytiva. cytivalifesciences.com/insights/surface-plasmon-resonance. Accessed September 9, 2025.
- Nguyen HH, Park J, Kang S, Kim M. Surface Plasmon Resonance: A Versatile Technique for Biosensor Applications. Sensors. 2015;15(5):10481-10510. doi:10.3390/s150510481
- Huber S, Casagrande F, Hug MN, et al. SPR-based fragment screening with neurotensin receptor 1 generates novel small molecule ligands. PLoS ONE. 2017;12(5):e0175842. doi:10.1371/journal.pone.0175842
- Heinrich T, Seenisamy J, Emmanuvel L, et al. Fragment-based discovery of new highly substituted 1H-pyrrolo[2,3-b]- and 3H-imidazolo[4,5-b]-pyridines as focal adhesion kinase inhibitors. J Med Chem. 2013;56(3):1160-1170. doi:10.1021/jm3016014
- Li XP, Harijan RK, Cao B, et al. Synthesis and Structural Characterization of Ricin Inhibitors Targeting Ribosome Binding Using Fragment-Based Methods and Structure-Based Design. J Med Chem. 2021;64(20):15334-15348. doi:10.1021/acs.jmedchem.1c01370
- Luttens A, Gullberg H, Abdurakhmanov E, et al. Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses. J Am Chem Soc. 2022;144(7):2905-2920. doi:10.1021/jacs.1c08402
- FitzGerald EA, Vagrys D, Opassi G, et al. Multiplexed experimental strategies for fragment library screening against challenging drug targets using SPR biosensors. SLAS Discov Adv Life Sci R D. 2024;29(1):40-51. doi:10.1016/j.slasd.2023.09.001
- Kim H, Weidner N, Ronin C, et al. Evaluating the affinity and kinetics of small molecule glycomimetics for human and mouse galectin-3 using surface plasmon resonance. SLAS Discov. 2023;28(5):233-239. doi:10.1016/j.slasd.2023.03.005
- Uitdehaag JCM, de Man J, Willemsen-Seegers N, et al. Target Residence Time-Guided Optimization on TTK Kinase Results in Inhibitors with Potent Anti-Proliferative Activity. J Mol Biol. 2017;429(14):2211-2230. doi:10.1016/j.jmb.2017.05.014
- Thinn AMM, Wang W, Chen Q. Competitive SPR chaser assay to study biomolecular interactions with very slow binding dissociation rate constant. Anal Biochem. 2025;696:115679. doi:10.1016/j.ab.2024.115679
- Quinn JG, Pitts KE, Steffek M, Mulvihill MM. Determination of Affinity and Residence Time of Potent Drug-Target Complexes by Label-free Biosensing. J Med Chem. 2018;61(12):5154-5161. doi:10.1021/acs.jmedchem.7b01829
- Zhukov A, Andrews SP, Errey JC, et al. Biophysical Mapping of the Adenosine A2A Receptor. J Med Chem. 2011;54(13):4312-4323. doi:10.1021/jm2003798
- Bobone S, Storti C, Fulci C, et al. Fluorescent Labeling Can Significantly Perturb Measured Binding Affinity and Selectivity of Peptide-Protein Interactions. J Phys Chem Lett. 2024;15(40):10252-10257. doi:10.1021/acs.jpclett.4c01767
- Livnat Levanon N, Vigonsky E, Lewinson O. Real Time Measurements of Membrane Protein:Receptor Interactions Using Surface Plasmon Resonance (SPR). J Vis Exp JoVE. 2014;(93):51937. doi:10.3791/51937
- Schneider CS, Bhargav AG, Perez JG, et al. Surface plasmon resonance as a high throughput method to evaluate specific and non-specific binding of nanotherapeutics. J Control Release Off J Control Release Soc. 2015;219:331-344. doi:10.1016/j.jconrel.2015.09.048
- Machine learning drives smarter SPR analysis. Selectscience.net. Published 2025. https://www.selectscience.net/article/machine-learning-drives-smarter-spr-analysis
- Navratilova I, Dioszegi M, Myszka DG. Analyzing ligand and small molecule binding activity of solubilized GPCRs using biosensor technology. Anal Biochem. 2006;355(1):132-139. doi:10.1016/j.ab.2006.04.021