By Sheffali Dash, Michelle Sabourin, Jichu Li, and Paula Ravnikar
Introduction
Cell line development is typically on the critical path of drug development. It’s a complex, labor intensive, and multistep process. Our cell line development platform uses the GOCHO™ host cell line to minimize the risk and timeline to develop high-performing clones suitable for manufacturing. However, multiple minipools and/or clones are often lost in the expansion process during the transition from static to suspension culture. In this study, we explored an alternative approach to reduce the losses of minipools and clones to improve the overall process result. In the cell line development workflow, minipool selection is initiated 48 h after transfection. Through a series of expansion steps, minipools or clones are expanded to progressively larger volumes and productivity is ranked. We investigated the parameters to use for optimal expansion and for timing the transition from static to shaking culture. With the new workflow described, we improved the overall timeline by ~ 2 weeks and readily obtained multiple clones producing > 4 g/L in a fed-batch culture. This new workflow is more robust than our previous version and increases our chances of delivering highly productive clones.
New workflow for minipool selection and expansion showed higher recovery
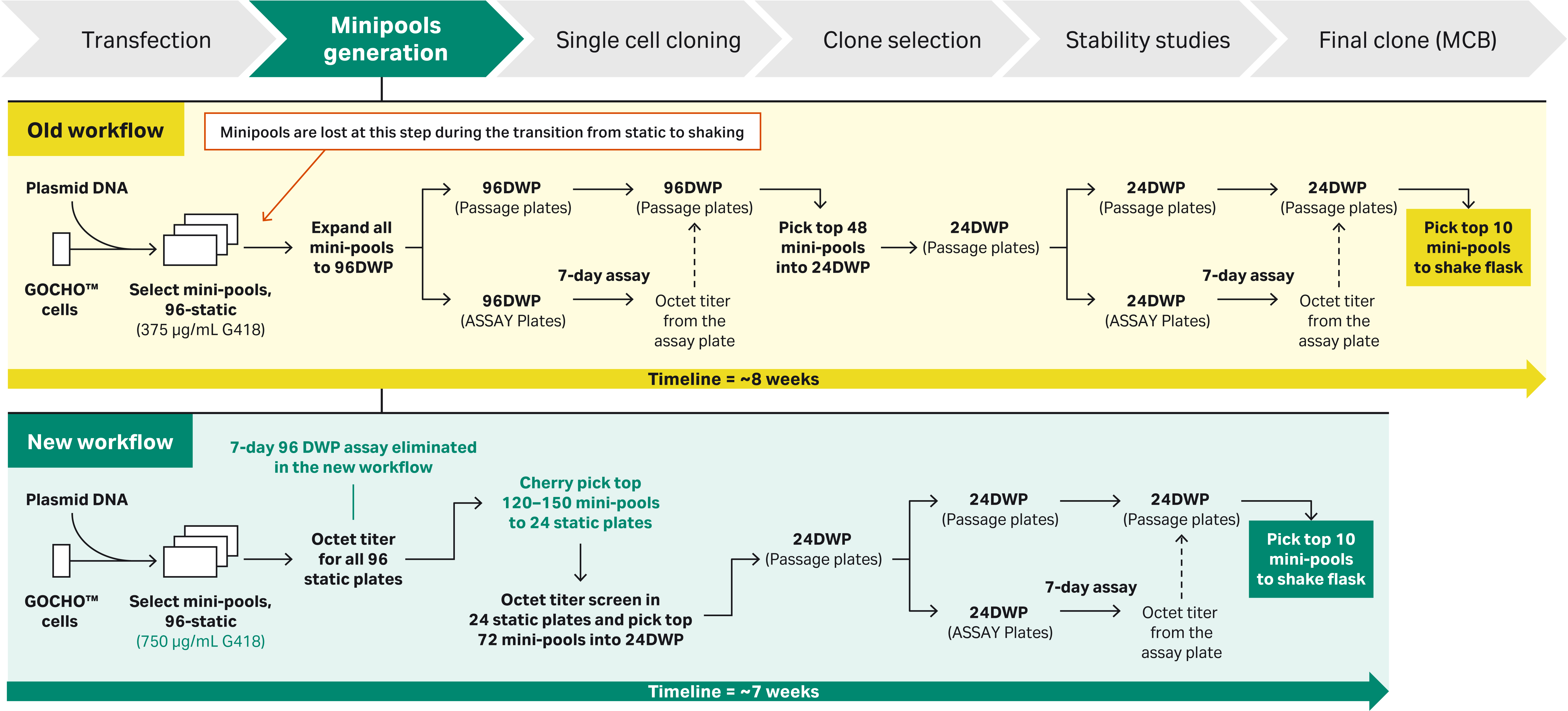
Fig 1. CLD workflow for minipools selection process: a) Old workflow for minipool generation using 96 DWPs. b) New workflow to minimize loss of minipools during expansion.
Note: The new workflow for clone expansion is similar but during expansion, clones are picked from different minipools to ensure heterogeneity.
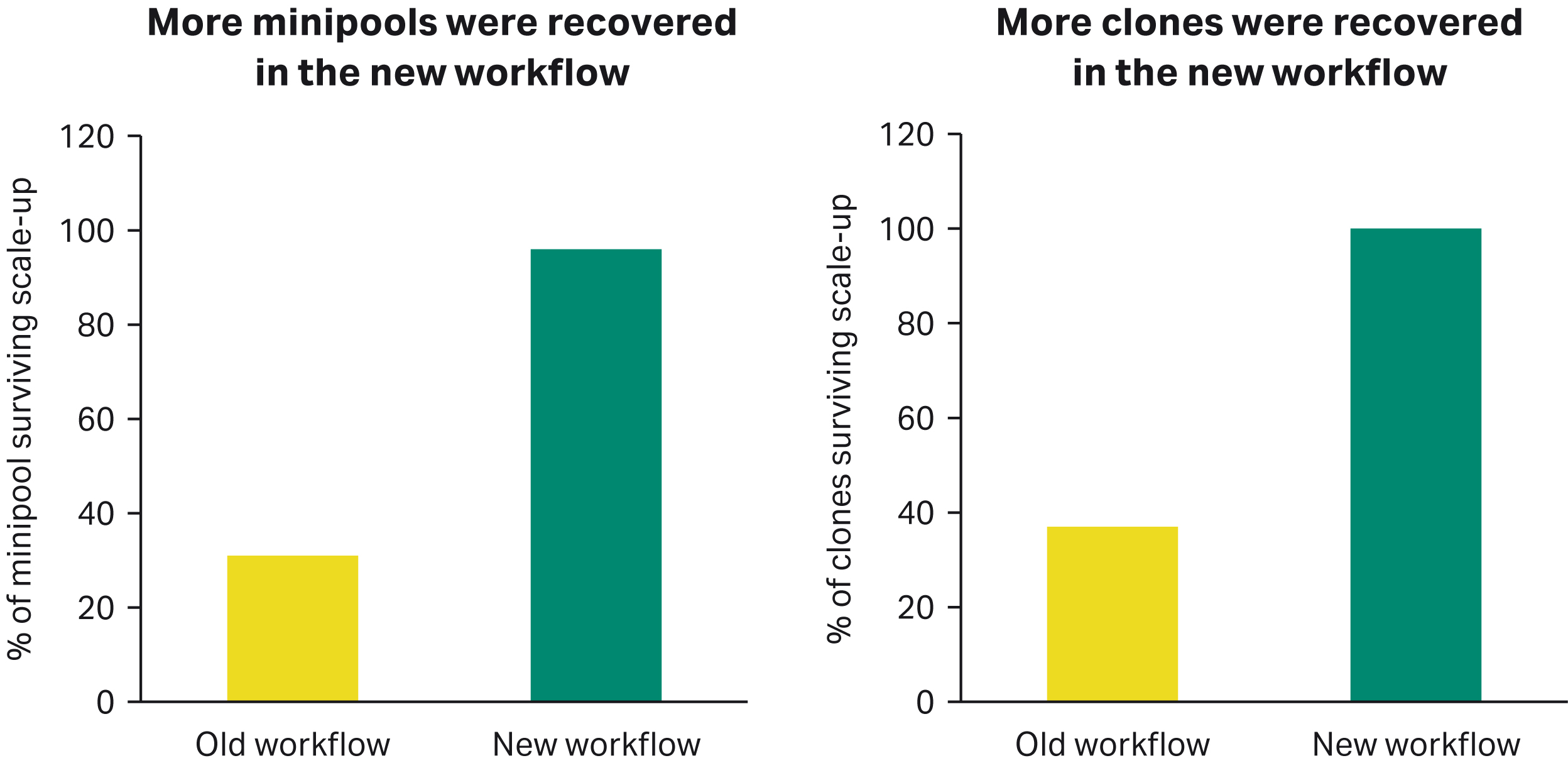
Higher recovery of minipools and clones in the new workflow
More minipools and clones were recovered in the new workflow at the expansion stage from 96-well static plates (old workflow scaled to 96 deep-well plates [DWP], and the new workflow scaled to 24-well static plates).
(A) (B)
Fig 2. Percent survival of minipools (A) and clones (B) during the expansion from 96-well static plates.
Top 10 minipools selected for single-cell cloning
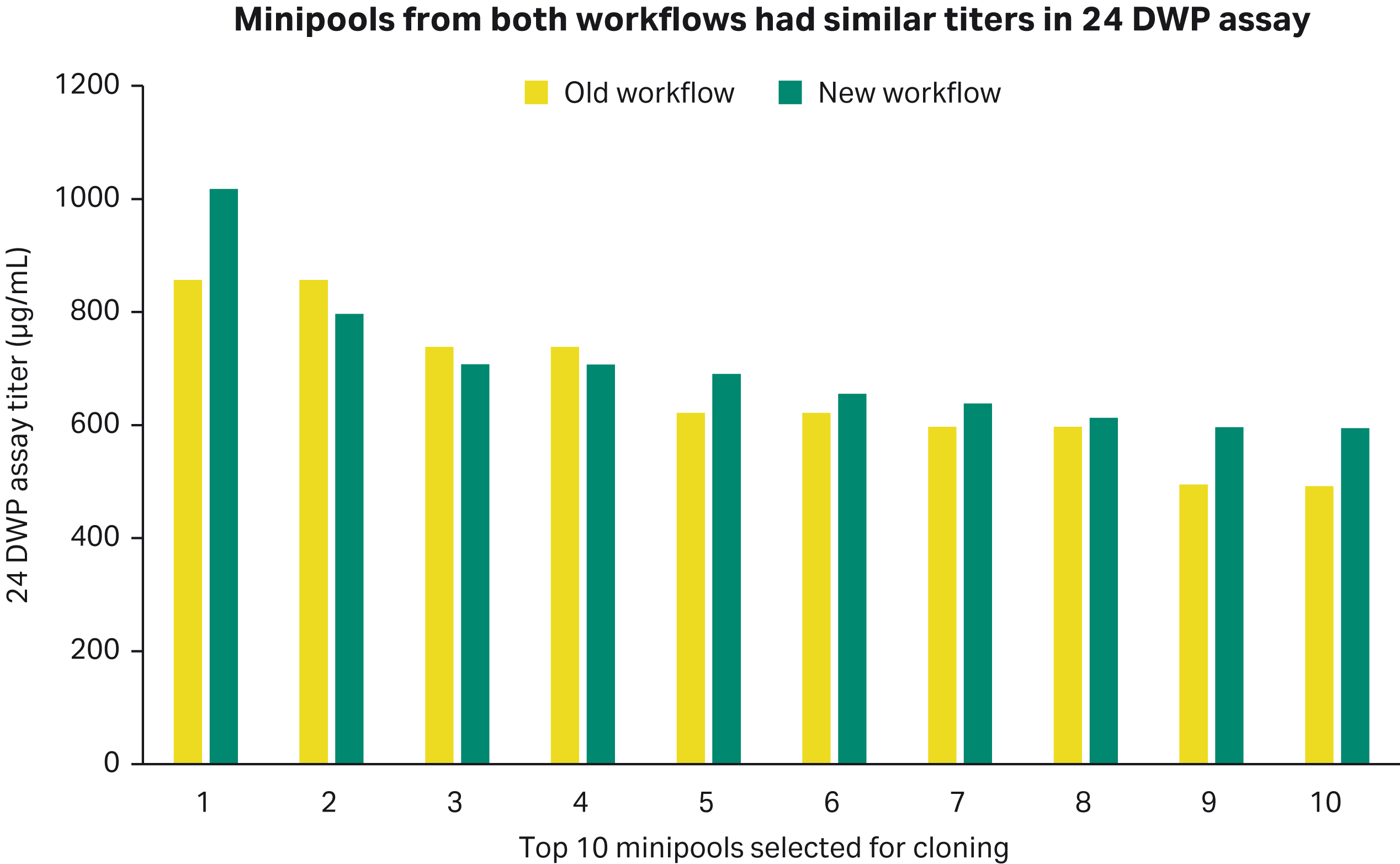
The minipools titers in the 24 DWP 7-day assay looked similar in both workflows. The minipools shown below were cloned by depositing 1 cell per well into 96-well static plates using single-cell printer™ from Cytena™.
Fig 3. 24 deep-well plate assay titer of minipools generated by both workflows.
New workflow resulted in more productive clones
Top clones were analyzed in a 14-day fed-batch assay
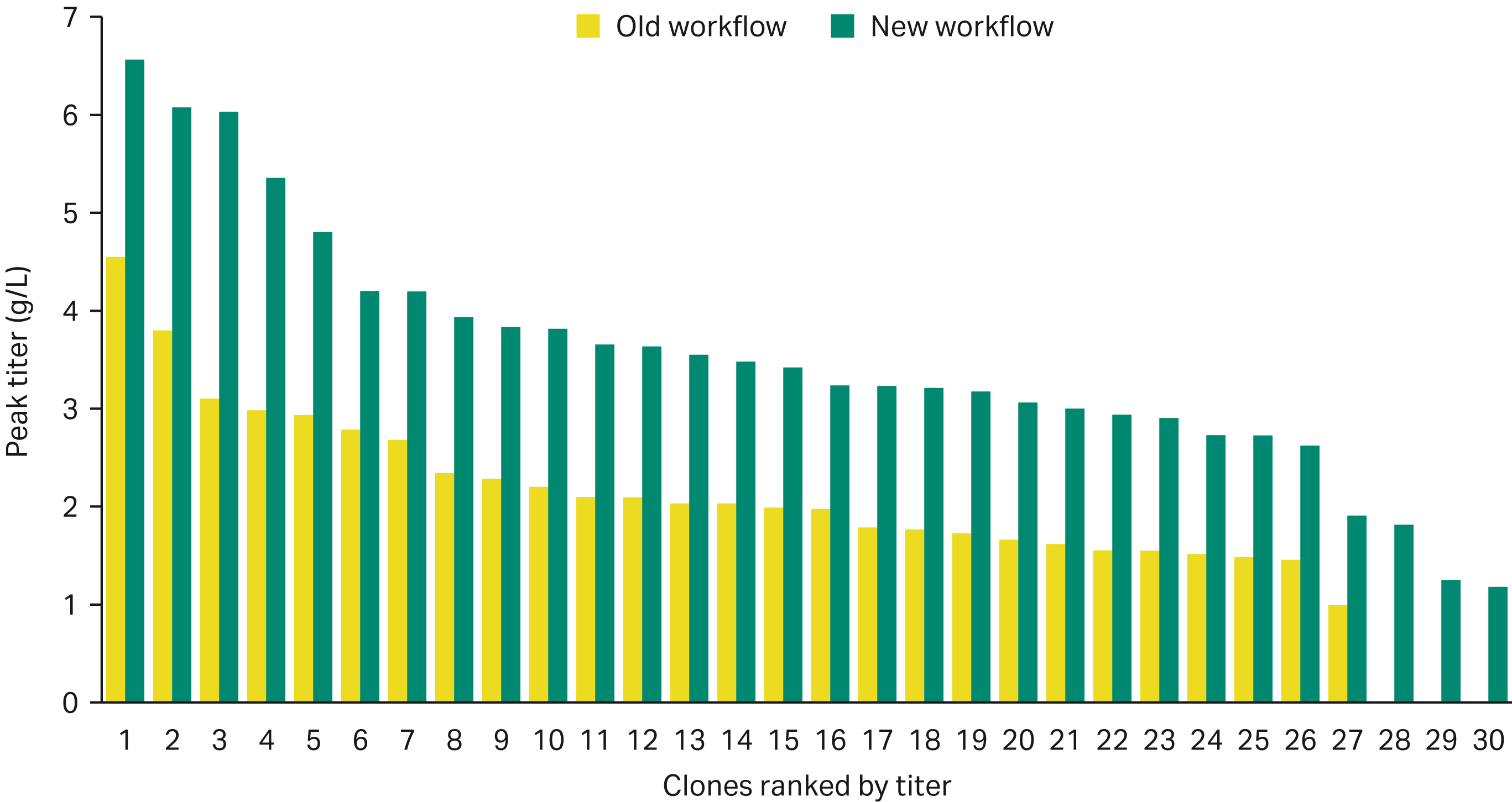
The resulting clones from the new workflow were more productive than the clones obtained from the old workflow. This could be due to higher recovery of clones during the expansion process. Selection with higher levels of G418 may have contributed to this outcome.
Fig 4. Higher producing clones were obtained from the new workflow with three clones making > 6 g/L in a 14-day fed-batch assay.
| Titer |
Old workflow (number of clones) |
New workflow (number of clones) |
| 2-3 g/L | 11 | 6 |
| 3-4 g/L | 2 | 13 |
| 4-5 g/L | 1 | 3 |
| 5-6 g/L | 0 | 1 |
| >6 g/L | 0 | 3 |
Table 1. Comparison of peak clone titers in a 14-day fed-batch assay using both workflows.
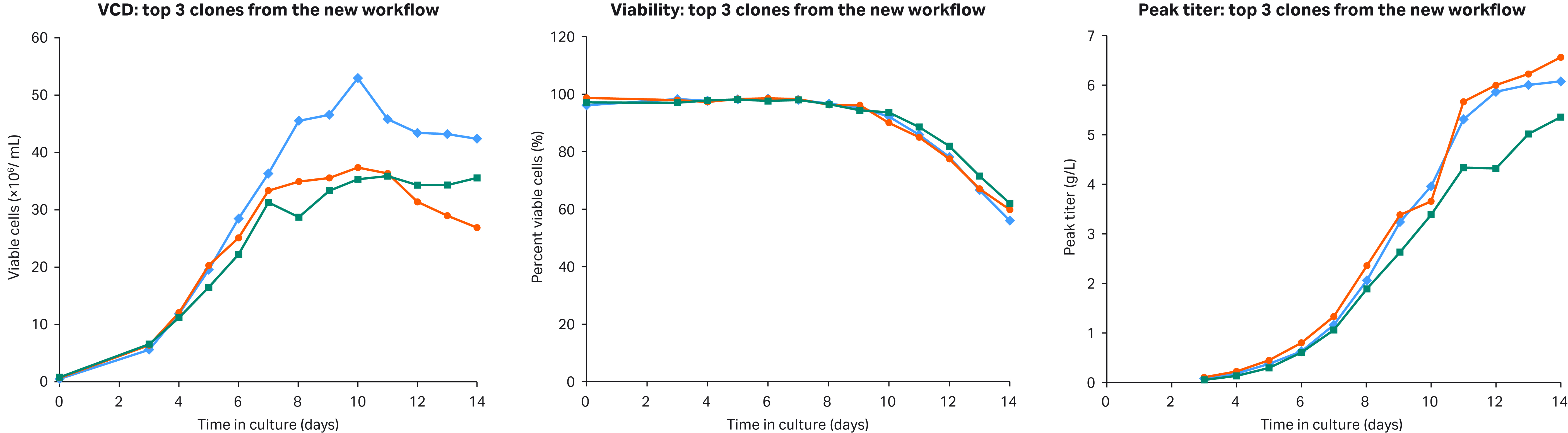
(A) (B) (C)
Fig 5. Shown above are the VCD (A), Viability (B), and titer (C) profiles for the top three clones obtained from the new workflow.
Future work
- Confirm robustness of the new workflow with additional molecules.
- The long-term stability of the clones obtained from the new workflow is being evaluated and will be compared with the stability obtained with the old workflow.
Conclusions
- We obtained multiple clones producing > 4 g/L in a fed-batch culture, which may be further optimized by process development. Three clones derived from two originating minipools produced > 6 g/L.
- In the new workflow described, we improved the overall timeline by ~ 2 weeks by eliminating the 7-day assay in 96 deep-well plates at both the selection step and the cloning step.
- The new process is more robust as a higher percentage of minipools and clones were recovered from the expansion process resulting in higher producing clones.
Learn more about cell line development services and our GOCHO™ platform.