Introduction
In protein research, going from idea to pure characterized protein takes multiple steps and techniques. When streamlined, this process provides fast results, which help you meet publishing deadlines and make critical project decisions.
Most scientists exploring proteins and developing specific target biomolecules use steps including cell culture, sample preparation, purification, and analysis. This requires understanding of the principles behind many different tools. Learning these steps takes time that scientists could use to generate data and progress research. If you’re short on time, setting up a strategy for complete protein development and production workflow can seem overwhelming.
Cytiva scientists Jon Lundqvist, Jinyu Zou, and Anna Moberg each have more than two decades of experience in protein research and Biacore™ SPR. This article covers their expert guidance, insight, and tips to help you make fast progress in protein research.
What are proteins?
Proteins are large biomolecules made up of amino acids. Proteins differ from each other by the sequence of amino acids and their specific 3D structure that determines activity. Proteins are responsible for every major cellular activity within organisms including:
- Metabolic reactions
- DNA replication
- Stimuli response
- Cell and organism structure
- Transporting molecules
Natural protein production in cells
Protein production is a vital biological process that occurs in the ribosome of a cell. Polypeptide chains are formed from various amounts of 20 different amino acids.
Steps in gene expression
Traditionally, scientists design DNA templates using RNA isolation from relevant host cells (e.g., mouse hepatocytes) followed by cDNA synthesis and polymerase chain reaction (PCR) amplification of the appropriate gene. The cloning strategy is designed within the PCR primers and allows cloning of the PCR insert into a suitable DNA vector to generate a protein expression plasmid.
Nowadays, synthetically made gene sequences are a time- and cost-effective alternative to traditional cloning. Vendors can codon-optimize synthetic genes for the intended expression host cell, increasing the likelihood of obtaining a high expression construct. Most vendors in the synthetic gene space also offer cloning services and can supply ready-to-use protein expression plasmids.
Recombinant protein production
Recombinant proteins (with or without introduced tags in the construct) are expressed in a protein host cell system (e.g., mammalian, prokaryotic, insect, yeast, etc.) through transformation and transfection with the expression plasmid. Larger amounts of expression plasmids are needed to obtain high levels of protein expression in certain systems, especially mammalian cells. To obtain these amounts, you can propagate the expression plasmid in suitable E. coli strains before plasmid purification and before transfecting the plasmid into the expression host cells.
Tags can be used to simplify identification, purification, and analysis of expressed proteins. The most common tag is histidine (His-tag) because it’s small and can remain on the protein after purification.
While His-tag is a good place to start, sometimes you’ll want to test one or more alternative tags like GST-tag, Strep-tag II, or MBP-tag. The tag needs to be soluble and attach to either the N- or C-terminal side on the target protein to make it accessible for detection and binding to the purification resin. If you need to remove the tag after purification, a cleavage site can be created in the tagged protein vector.
Steps in recombinant protein production
You might want a one-size-fits-all approach to your protocols, but this can cost time in the long run. You need to adjust protocols to fit specific proteins, and add, change, or remove steps for specific targets. One-protocol-fits-all thinking can lead to low expression, poor yield, and time-consuming troubleshooting.
Before setting up the protocol, set your goals and objectives.
What is the purpose of your research? Will your protein be used to determine structure and function or for therapeutic and diagnostic use?
How much information do you have on the identity and function of the target protein?
What analysis methods will you use?
Do you need post-translational modifications (PTM), such as glycosylation, for protein functionality? Will that influence the expression system you choose?
Clear goals help you decide which expression system to use, the purity and yield to aim for, and the amount and quality of sample you need to conduct multiple analyses. Setting goals also helps you choose which analysis methods to use.
When it comes to choosing equipment and consumables, do some extra research. Different proteins have different properties. Reusing protocols and existing equipment might not provide the outcomes you want. Search the literature in your field and ask around for more insight. Your colleagues have a wealth of insight depending on the size of your lab.
Cell culture
Selecting an expression system
There are several factors to think about when selecting an expression system. If you haven’t already, fill out the goals and objectives for your research we outlined earlier. You can select the expression system based on the purpose of purified protein, the amount, purity level, and requirement for different variants.
You will need to evaluate whether to use a prokaryotic or eukaryotic expression system. E. coli is the go-to-platform for many recombinant proteins. It has well-known genetics, a high transformation efficiency, and it’s easy, fast, and inexpensive to cultivate. However, if your protein doesn’t express well in E. coli, then an insect or mammalian system can be more efficient and cost-effective.
In addition to your larger goals, keep in mind some practical considerations:
Structural complexity: Bacterial proteins and eukaryotic proteins with a simple tertiary structure are good candidates for expression in bacterial systems. Eukaryotic proteins with a more complex tertiary structure are better candidates for expression in mammalian or insect cell systems that can support protein folding and post-translational modifications.
Post-translational modifications (PTMs): PTMs can be critical for a protein’s folding, function, and localization. These modifications include phosphorylation, glycosylation, prenylation, disulfide bonds, protease cleavage, and more. Mammalian cells have the most sophisticated PTM pathways.
Solubility: Eukaryotic proteins expressed in bacteria can be biologically inactive and localized to insoluble inclusion bodies because PTM systems are lacking, disulfide bond formation is difficult, and bacterial cells don’t have the same chaperone networks as eukaryotic cells to assist with protein folding. In some cases, you can purify your protein of interest from inclusion bodies using solubilization and refolding steps. If that doesn’t work, then you might want to use expression in a eukaryotic system.
Cellular localization: A bacterial, insect, or mammalian cell system can be a viable option for recombinant expression of an intracellular protein. The expression of extracellular and/or membrane-bound proteins present additional challenges – in many cases, the PTM and intracellular pathways in eukaryotes make the insect and mammalian systems the go-to option for these proteins.
Toxicity: Some exogenous proteins’ expression can be toxic to a host cell. To overcome this, you can use special expression vectors and modified E. coli strains. In mammalian cells, you can also consider using inducible systems (e.g., tetracycline-regulated on/off).
Cell culture conditions
To optimize protein expression, you should screen cell culture conditions in 24- to 48-well plates and manage 1 to 2 mL cultivations in parallel.
During the screening, conditions like temperature and additives vary. If the expression levels are lower than you expect, or if the target protein is not expressed, go back and look at factors like the tag system and cultivation conditions. Optimize them until you find the clone and expression levels that meet the goals you set.
Once you’ve chosen an expression system and the conditions to grow the host cell, you can scale up to larger cell culture volumes. Scientists commonly use cell culture volumes from 0.1 to 20 L in research scale and use shaker flasks when working with E. coli.
For higher expression levels and mammalian cells, you can use a fermenter to better control conditions like temperature, oxygen, glucose levels, and pH. Exact conditions vary depending on cell type, but all cultures have a vessel and medium that support cell growth by supplying hormones, growth factors, and nutrients. To prolong mammalian cell viability and growth, use specific medium feeds in conjunction with chemically defined media, or supplement classic formulations with serum.
The physicochemical environment (pH, osmolality, etc.) is also critical to successful cell culture. You can maintain it by using process liquids and buffers.
Sample prep
Throughout sample preparation, protein purification, and analysis, you need to avoid protein damage and maintain protein activity.
Some proteins are more sensitive than others. Understanding your protein helps you determine which conditions to use. Some proteins, like secreted proteins, have lower expression levels. When expression levels are low It’s even more important to maintain high yields in each step.
Sample preparation depends on protein type. Membrane proteins, protein complexes, and secreted proteins have different sample preparation protocols. For intracellular proteins, scientists lyse the cells to release the target. This can be done by sonication, homogenization, freezing and thawing, or chemical agents.
For extracellular proteins obtained with mammalian or insect cells, you can directly apply cell harvest. Scientists often perform cell harvest using centrifugation. In most applications, scientists transfer the supernatant to protein purification containing the target protein or analyze it directly to perform identification and determine activity.
If needed, filtration can be used to remove particles after centrifugation or before chromatography to protect columns. Desalting at laboratory scale is a well-proven, simple, and fast method that removes low molecular weight impurities and transfers the sample into required buffer in a single step.
Desalting or buffer exchange can be performed prior to a chromatography step or between purification steps for sample conditioning.
Protein purification
In protein research, scientists often use two purification steps: Affinity chromatography and size exclusion chromatography(SEC). If you need high purity, add an additional intermediate step of ion exchange or hydrophobic interaction chromatography. However, try to use as few steps as possible, adding steps decreases overall protein yield.
Tagged recombinant proteins are usually straightforward to purify. Use an affinity resin that’s suitable for your tag system and use the protein’s natural conditions to avoid precipitation and degradation.
You might need to remove the tag after purification. After tag cleavage, separate tag-free target protein from the tag and tag-containing target protein. You can also use on-column cleavage. Add protease to the column with the target protein bound to enable one-step cleavage and removal.
Scientists often use affinity chromatography for antibody purification. Affinity chromatography uses the high affinity and specificity of protein A and protein G for the Fc-region of IgG from many species. Protein L, which binds to kappa light chains, is another ligand you can use to purify antibody fragments, IgG, and other antibodies from a wide range of eukaryotic species. For higher purity, scientists often use SEC as a second step.
You’ll need more effort when developing the protocols for nontagged proteins. Experiment planning helps you decide which conditions to test, as well as what to optimize to achieve your desired purity and yield.
Scientists commonly use gravity columns in affinity steps and occasionally use peristatic pumps in other chromatography steps. A protein purification system, often in combination with a prepacked column, delivers significantly more control so you can obtain more detailed and consistent information on your target protein and any impurities. It also provides better column protection.
Sign up for Purify club for more insights into protein purification.
Protein analysis – identification and characterization
Proteins are diverse in size, structure, and biochemical properties. Scientists use these differences to characterize isolated proteins. You can use multiple techniques to confirm identity and binding activity during protein expression, production, in between steps, and within the purified protein.
A lot of techniques and methods are common in analysis and characterization. UV, SDS-PAGE, and SEC are popular choices for identifying which fractions contain the target protein. Assays for specific proteins like enzymatic assays are available and require a substrate specific to the target protein.
After purification, you can determine structures, binding activities, stability, and more. That’s often done using mass spectrometry in combination with high-performance liquid chromatography (HPLC) columns, amino acid sequencing, X-ray crystallography, cryo electron microscopy (cryo-EM), surface plasmon resonance (SPR), western blotting, circular dichroism, and nuclear magnetic resonance (NMR) spectroscopy.
Western blotting
Western blotting is one of the most established and popular techniques for quantifying and identifying proteins. The method builds an antibody:protein complex. Scientists bind the antibodies to membrane-immobilized proteins and detect the bound antibody with a detection method like chemiluminescence or fluorescence. A sample can be a complex protein mixture like a cell or tissue extract, but it can also be a sample of purified proteins, like a fraction from a purification run.
The Western blot workflow has several steps before analysis:
Scientists apply gel electrophoresis to the sample for protein separation before immobilizing the proteins on a membrane, (e.g., nitrocellulose or PVDF) following electrotransfer from the gel.
Scientists then incubate the membrane with a primary antibody that specifically binds to the protein of interest, wash to remove unbound antibodies, and conjugate a secondary antibody to an enzyme, a fluorophore, or an isotope for detection. The detected signal from the protein:antibody:antibody complex is proportional to the amount of protein on the membrane.
Traditional total protein staining methods often limit the dynamic range, typically by one or two orders of magnitude for silver and Coomassie staining. In contrast, fluorescence and chemiluminescence detection has a much broader detection window and is quick, easy to use, and highly sensitive.
You can use a CCD-based imager to visualize the blot in both chemiluminescence and fluorescence ranges with high sensitivity and resolution.
How can SPR help with protein analysis?
Understanding the nature of molecular interactions is critical in all areas of life sciences. Label-free interaction analysis generates data-rich content that can help enhance our understanding of interactions between almost any type of biologically relevant molecules.
Do the potential interactants bind to each other?
How specific is the interaction?
How strong is the binding (i.e., affinity)?
How fast is the binding?
What are the effects of temperature?
How much interactant is there in the sample?
Biacore™ SPR systems provide real-time analysis of events that are driven and regulated through dynamic interactions between key molecules.
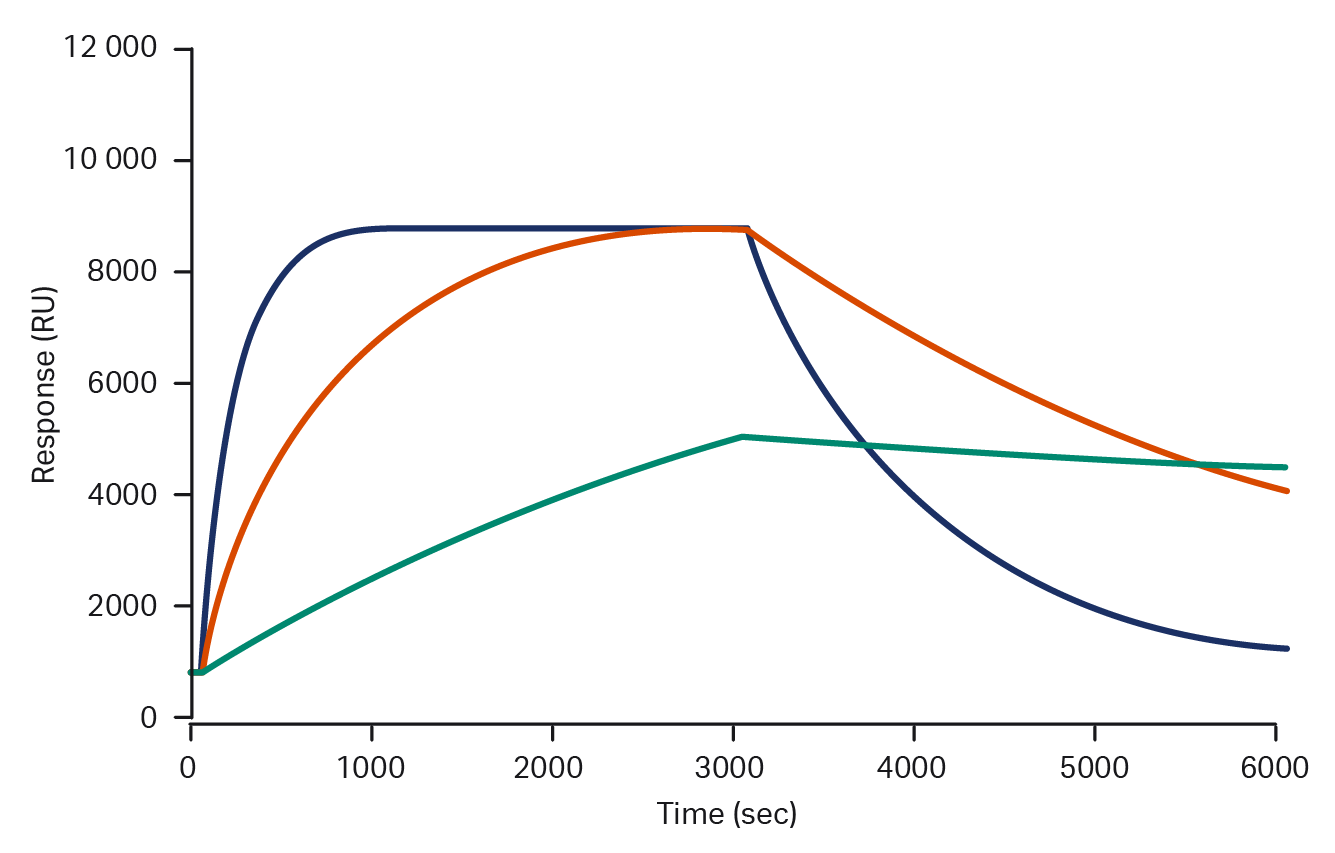
Biological processes are real-time events that are driven and regulated through dynamic interactions between key molecules. End-point techniques like ELISA can only offer a snapshot and provide basic information like overall binding strength. Affinity depends on the ratio of on- and off-rates, so equal affinity interactions can have very different kinetic properties. Biacore™ systems enable real-time analysis that can provide the data needed to discriminate these crucial differences.
Sensorgrams showing three molecules with identical affinities, but kinetic profiles differing by several orders of magnitude. These differences are not visible with end-point analyses.
Biacore™ sensor chips support analysis of a wide range of interactions. You can use kits and ready-made consumables to save time while maintaining consistent capture of molecules e.g., antibodies and common tags.
We recommend using predefined Biacore™ system run methods with application-relevant settings. Combine them with evaluation software for a good data overview that shortens time to results.
Sign up for Biacore™ system newsletters for more insights in protein purification and SPR interaction analysis.
Challenges and advances in protein production and characterization
|
Challenges |
Advancements |
|
Complex protein formats Producing bispecific antibodies and other multi-chain proteins is technically demanding. Mispaired chains can result in non-functional products, complicating downstream analysis.
|
High-throughput screening tools and platforms like Biacore™ 8K+ system provide simultaneous analysis of multiple interactions to identify correctly paired proteins more efficiently. |
|
Product-related impurities Aggregates, homodimers, and half-mers often mimic the target protein structurally but lack functionality. These impurities are hard to detect with conventional assays and require sensitive tools like SPR.
|
SPR-based systems such as Biacore™ SPR instruments offer high sensitivity and specificity for precise detection and differentiation of functional proteins from impurities in crude samples.
|
|
Limited sample availability Early-stage research or rare proteins often yield low sample volumes, making it difficult to perform comprehensive characterization without highly sensitive platforms.
|
Biacore™ systems are optimized for low-volume, high-sensitivity assays, to provide reliable kinetic and affinity measurements even with minimal sample input. |
|
Time-intensive workflows Traditional protein production and purification can take weeks, delay assay readiness, and slow down development timelines.
|
Biacore™ systems can be integrated with platforms for rapid protein expression for a streamlined workflow from production to characterization. This reduces turnaround time and accelerates decision-making in early-stage development. |
Summary and outlook
As the biologics landscape evolves toward more complex formats, such as bispecifics, antibody-drug conjugates, and PROTACs, (links to upcoming whitepapers later) the demand for precision, speed, and scalability in production and characterization intensifies. At the same time, the rise of AI-optimized protein design accelerates the pace of discovery, which requires platforms that can keep up with rapid iteration and validation cycles.
Integrated solutions like Biacore™ SPR systems, when paired with rapid protein expression platforms, are well-positioned to meet these demands. They provide seamless transitions from design to functional analysis, supporting both exploratory research and translational development. Continued innovation in automation, data analytics, and assay sensitivity will be key to unlocking the full potential of next-generation biologics.
FAQs
What are the different types of protein production?
Common protein production systems are derived from bacteria, yeast, insect, or mammalian cells.
What are the major steps in protein production?
The major steps in protein production include:
- Cell culture
- Sample preparation
- Protein purification
- Protein analysis: Identification and characterization
How does SPR work for studying protein interactions?
Biacore™ SPR systems deliver real-time analysis of events that are driven and regulated through dynamic interactions between key molecules. Affinity depends on the ratio of on- and off-rates, so equal affinity interactions can have very different kinetic properties. Biacore™ systems provide the data needed to discriminate between these crucial differences and are designed to generate high quality, information-rich data that answers key questions concerning the nature of binding — how strong, how fast, how much, and how specific — driving your project to conclusion.